Difference between revisions of "Projects/Diffusion/2007 Project Week Slicer 3 Whole Brain Seeding"
From NAMIC Wiki
| (3 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
{| | {| | ||
|[[Image:ProjectWeek-2007.png|thumb|320px|Return to [[2007_Programming/Project_Week_MIT|Project Week Main Page]] ]] | |[[Image:ProjectWeek-2007.png|thumb|320px|Return to [[2007_Programming/Project_Week_MIT|Project Week Main Page]] ]] | ||
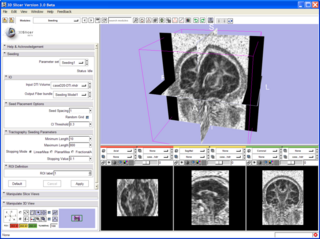
| − | |[[Image: | + | |[[Image:SeedingInterface.png|thumb|320px|Slicer 3 tractography display.]] |
|} | |} | ||
| Line 23: | Line 23: | ||
Our approach is to port the Streamline techniques used in Slicer 2 as a Command Line Module. | Our approach is to port the Streamline techniques used in Slicer 2 as a Command Line Module. | ||
We plan on looking at a optimal memory representation of a full brain tractography set for optimal querying and retrival of specific fibers according to spatial locations. | We plan on looking at a optimal memory representation of a full brain tractography set for optimal querying and retrival of specific fibers according to spatial locations. | ||
| + | |||
| + | </div> | ||
<div style="width: 40%; float: left;"> | <div style="width: 40%; float: left;"> | ||
| − | |||
| − | |||
<h1>Progress</h1> | <h1>Progress</h1> | ||
| − | Software for the fiber tracking has been ported. | + | * Software for the fiber tracking has been ported and interfaces has been updated to VTK 5. |
| + | * The command line module has been developed and fiber mrml bundles have been incorporated to the XML description (joint collaboration with Casey Goodlett). | ||
| + | * Collaboration with parties involved in Diffusion Imaging to facilitate the incorporation of diffusion tools. | ||
| + | * Design decision has been made to incorporate basic display logic into DisplayMRMLNodes to meet needs across projects and improve performance. | ||
</div> | </div> | ||
Latest revision as of 14:15, 29 June 2007
Home < Projects < Diffusion < 2007 Project Week Slicer 3 Whole Brain Seeding Return to Project Week Main Page |
Key Investigators
- BWH: Raul San Jose, Lauren O'Donnell, Alex Yarmarkovich
Objective
Deployment of whole brain seeding: data structure, fiber tracking and fast query mechanism to support editing operation and ROI-based fiber selection.
Approach, Plan
Our approach is to port the Streamline techniques used in Slicer 2 as a Command Line Module. We plan on looking at a optimal memory representation of a full brain tractography set for optimal querying and retrival of specific fibers according to spatial locations.
Progress
- Software for the fiber tracking has been ported and interfaces has been updated to VTK 5.
- The command line module has been developed and fiber mrml bundles have been incorporated to the XML description (joint collaboration with Casey Goodlett).
- Collaboration with parties involved in Diffusion Imaging to facilitate the incorporation of diffusion tools.
- Design decision has been made to incorporate basic display logic into DisplayMRMLNodes to meet needs across projects and improve performance.