Difference between revisions of "2017 Winter Project Week/MeningiomaSegmentation"
From NAMIC Wiki
m |
|||
| (3 intermediate revisions by the same user not shown) | |||
| Line 8: | Line 8: | ||
==Key Investigators== | ==Key Investigators== | ||
<!-- Add a bulleted list of investigators and their institutions here --> | <!-- Add a bulleted list of investigators and their institutions here --> | ||
| + | * Jakub Kaczmarzyk, MIT | ||
* Satrajit Ghosh, MIT | * Satrajit Ghosh, MIT | ||
* Omar Arnaout, Brigham and Women's Hospital | * Omar Arnaout, Brigham and Women's Hospital | ||
| Line 36: | Line 37: | ||
* Continue testing existing segmentation methods. | * Continue testing existing segmentation methods. | ||
* Try Slicer in a Nipype workflow. | * Try Slicer in a Nipype workflow. | ||
| − | * Apply manifold learning | + | * Apply feature engineering techniques, like manifold learning. |
* Get more data, and potentially train a neural network. | * Get more data, and potentially train a neural network. | ||
|} | |} | ||
| + | |||
| + | |||
| + | ==Examples== | ||
| + | {| class="wikitable" | ||
| + | | | ||
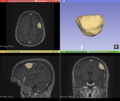
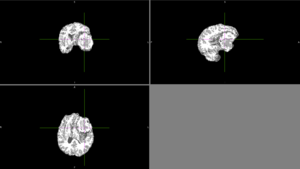
| + | [[File:Case_001_ants_brain_failure.png|thumbnail|Sometimes, ANTs failed to remove parts of the skull close to the tumor or wrongly removed part of the brain.]] | ||
| + | [[File:Case_052_ants_brain.png|thumbnail|Other times, ANTs extracted the brain well.]] | ||
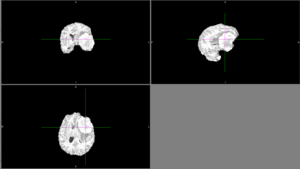
| + | [[File:Case_052_ants_brain_seg.png|thumbnail|Automatic segmentation with ANTs usually could not distinguish the tumor from the rest of the brain.]] | ||
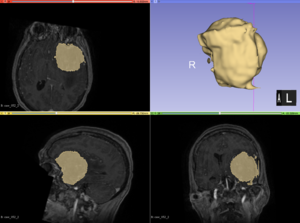
| + | [[File:Case_052_2_slicer_seg.png|thumbnail|Semi-automatic segmentation with Slicer was relatively successful. Sometimes the segmentation would bleed outside of the tumor into voxels with similar intensities.]] | ||
| + | | | ||
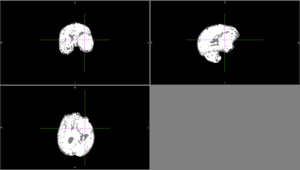
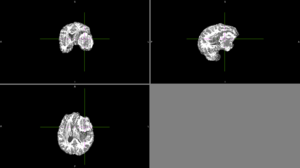
| + | We tried FSL's FAST with different numbers of classes. None of these methods could identify the entire tumor mass as one type of tissue in this scan. | ||
| + | [[File:Case_052_fast_3classes.png|thumbnail|FAST brain segmentation with 3 classes]] | ||
| + | [[File:Case_052_fast_4classes.png|thumbnail|FAST brain segmentation with 4 classes]] | ||
| + | [[File:Case_052_fast_5classes.png|thumbnail|FAST brain segmentation with 5 classes]] | ||
| + | |} | ||
| + | |||
| + | |||
==Background and References== | ==Background and References== | ||
<!-- Use this space for information that may help people better understand your project, like links to papers, source code, or data --> | <!-- Use this space for information that may help people better understand your project, like links to papers, source code, or data --> | ||
MR images of meningiomas that will be used in this project are available at [http://openneu.ro/metasearch/ OpenNeu.ro]. | MR images of meningiomas that will be used in this project are available at [http://openneu.ro/metasearch/ OpenNeu.ro]. | ||
Latest revision as of 21:20, 13 January 2017
Home < 2017 Winter Project Week < MeningiomaSegmentationKey Investigators
- Jakub Kaczmarzyk, MIT
- Satrajit Ghosh, MIT
- Omar Arnaout, Brigham and Women's Hospital
Project Description
| Objective | Approach and Plan | Progress and Next Steps |
|---|---|---|
|
|
Progress
Next steps
|
Examples
|
|
We tried FSL's FAST with different numbers of classes. None of these methods could identify the entire tumor mass as one type of tissue in this scan. |
Background and References
MR images of meningiomas that will be used in this project are available at OpenNeu.ro.