Difference between revisions of "Linear Mixed-effects shape model to explore Huntington's Disease Data"
From NAMIC Wiki
| Line 33: | Line 33: | ||
* preliminary results look promising | * preliminary results look promising | ||
* ITKv4 module for mixed-effects model (Josh) available for Manasi to test | * ITKv4 module for mixed-effects model (Josh) available for Manasi to test | ||
| + | ** ITKv4 module tested on Linux system | ||
| + | ** Mixed-effects model for single component HD shapes (in progress) | ||
* Once tested, this module can be used by UIowa team on a larger cohort of cases | * Once tested, this module can be used by UIowa team on a larger cohort of cases | ||
Revision as of 13:45, 20 June 2013
Home < Linear Mixed-effects shape model to explore Huntington's Disease DataKey Investigators
- UIowa: Hans Johnson, Dave Welch
- Utah: Manasi Datar, Ross Whitaker
Objective
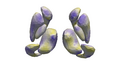
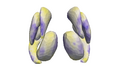
Make the linear mixed-effects shape model accessible for further exploration of Huntington's Disease Data
Approach, Plan
Meet with UIowa team to:
- give an overview of results from the linear mixed-effects shape model
- explain the ShapeWorks command line tools to optimze and analyze correspondences
- discuss next steps toward making these tools accessible to UIowa team, to facilitate further exploration of the Huntington's disease Data
Progress
- preliminary results look promising
- ITKv4 module for mixed-effects model (Josh) available for Manasi to test
- ITKv4 module tested on Linux system
- Mixed-effects model for single component HD shapes (in progress)
- Once tested, this module can be used by UIowa team on a larger cohort of cases
References
M Datar, P Muralidharan, A Kumar, S Gouttard, J Piven, G Gerig, RT Whitaker, PT Fletcher, Mixed-Effects Shape Models for Estimating Longitudinal Changes in Anatomy, STIA 2012