Difference between revisions of "2009 Summer Project Week Slicer3 Cortical Thickness Pipeline"
From NAMIC Wiki
(Created page with '__NOTOC__ <gallery> Image:PW2009-v3.png|Project Week Main Page Image:Arctic_Logo.png|[http://www.nitrc.org/projects/arctic ARCTIC: Automatic Regio...') |
|||
| Line 4: | Line 4: | ||
Image:Arctic_Logo.png|[http://www.nitrc.org/projects/arctic ARCTIC: Automatic Regional Cortical ThICkness] | Image:Arctic_Logo.png|[http://www.nitrc.org/projects/arctic ARCTIC: Automatic Regional Cortical ThICkness] | ||
Image:BrainDevelopment.jpg|Brain development | Image:BrainDevelopment.jpg|Brain development | ||
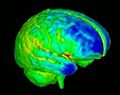
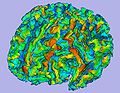
| − | Image: | + | Image:MeshBasedCortThick_CorticalThickness.jpg|Cortical thickness on white matter surface |
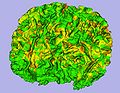
| − | Image: | + | Image:MeshBasedCortThick_SulcalDepth.jpg|Sucal depth on genus-zero cortical surface |
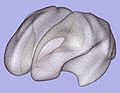
| − | Image: | + | Image:MeshBasedCortThick_Particles.jpg|Particles on inflated genus-zero cortical surface |
</gallery> | </gallery> | ||
Revision as of 21:20, 4 June 2009
Home < 2009 Summer Project Week Slicer3 Cortical Thickness PipelineKey Investigators
- UNC: Clement Vachet, Cedric Mathieu, Ipek Oguz, Marc Niethammer, Heather Cody Hazlett, Martin Styner
Objective
We would like to create end-to-end applications within Slicer3 allowing individual and group analysis of regional (voxel-based) and local (mesh-based) cortical thickness.
Such a workflow applied to the young brain (2-4 years old) is our goal in order to start a longitudinal study of early brain development in autism (UNC DBP).
See our Roadmap for more details.
Approach, Plan
Our approach is described in the following wiki pages:
Our plan for the project week is to:
- Improve ARCTIC's integration within Slicer3
- ARCTIC's availability using Slicer3's extension manager interface
- Cortical thickness overlaid on VTK surface (quality control using Slicer3 MRML scene)
- Future directions:
- Batch-processing within Slicer3 to perform group analysis
- Use of XNAT to directly download atlases needed for processing.
- Continue on the mesh-based cortical thickness pipeline.
- Particle correspondence individual testing
- Automated processing of small pediatric dataset
We are also interested in learning about batch-processing within Slicer3, XNAT integration for future improvment.
Progress
References
- I. Oguz, M. Niethammer, J. Cates, R. Whitaker, T. Fletcher, C. Vachet, and M. Styner, Cortical Correspondence with Probabilistic Fiber Connectivity, Information Processing in Medical Imaging, IPMI 2009, LNCS, in print.
- H.C. Hazlett, C. Vachet, C. Mathieu, M. Styner, J. Piven, Use of the Slicer3 Toolkit to Produce Regional Cortical Thickness Measurement of Pediatric MRI Data, presented at the 8th Annual International Meeting for Autism Research (IMFAR) Chicago, IL 2009.
- C. Mathieu, C. Vachet, H.C. Hazlett, G. Geric, J. Piven, and M. Styner, ARCTIC – Automatic Regional Cortical ThICkness Tool, UNC Radiology Research Day 2009 abstract
- C. Vachet, H.C. Hazlett, M. Niethammer, I. Oguz, J.Cates, R. Whitaker, J. Piven, M. Styner, Mesh-based Local Cortical Thickness Framework, UNC Radiology Research Day 2009 abstract