Difference between revisions of "2011 Summer Project Week Customizing EMSegmenter pipelines for brain lesions"
From NAMIC Wiki
Belhachemi (talk | contribs) |
Belhachemi (talk | contribs) |
||
| Line 47: | Line 47: | ||
<gallery perrow=1: widths=1250px : heights=400px> | <gallery perrow=1: widths=1250px : heights=400px> | ||
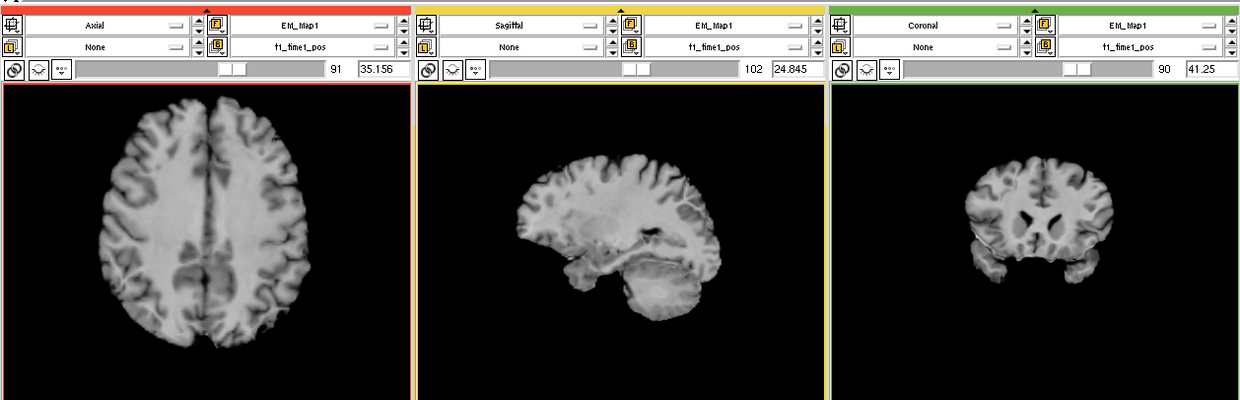
| − | Image:EMS LesionExp T1.png | + | Image:EMS LesionExp T1.png | T1 scan |
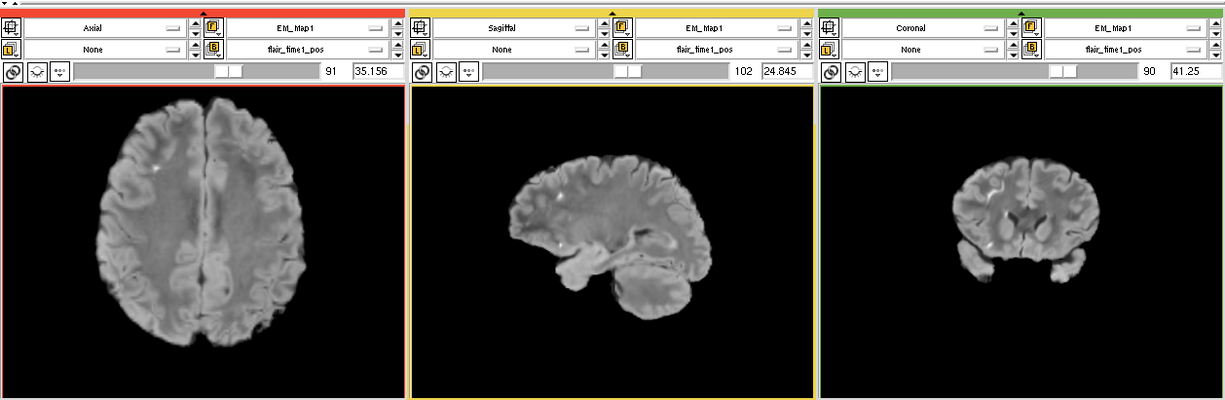
| − | Image:EMS LesionExp FLAIR.png | + | Image:EMS LesionExp FLAIR.png | FLAIR scan |
| − | |||
</gallery> | </gallery> | ||
Revision as of 16:57, 20 June 2011
Home < 2011 Summer Project Week Customizing EMSegmenter pipelines for brain lesions- Notfound.png
Interesting picture to be added...
Customizing EMSegmenter pipelines for brain lesions
Key Investigators
- University of Pennsylvania: Dominique Belhachemi
- University of Pennsylvania: Kilian Pohl
- Brigham and Women's Hospital: Alex Zaitsev
Objective
The goal is to segment white matter lesions based on T1 and FLAIR brain scans.
Approach, Plan
We want to use the EMSegmenter modle for this project. The plan is to create a two channel pipeline which includes pre-processing (e.g. registration) and applying the EM algorithm to perform the segmentation.
Progress
References
http://www.slicer.org/slicerWiki/index.php/EMSegmenter-Tasks
Delivery Mechanism
This work will be delivered to the NAMIC Kit as a
- Slicer Module
- Built-in:
- Extension -- commandline:
- Extension -- loadable: