2017 Winter Project Week/Population Based Image Imputation
From NAMIC Wiki
Home < 2017 Winter Project Week < Population Based Image Imputation
Key Investigators
- Adrian Dalca, MIT, MGH
- Katie Bouman, MIT
- Mert Sabuncu, MGH, Cornell
- Polina Golland, MIT
Project Description
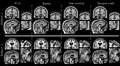
We developed a model for image imputation - or restoration - for clinical quality images where slice separation (e.g. 6mm) is significantly larger than slice resolution (e.g. 1mm^2). Our model captures statistical correlations within a collection of clinical images from a population of subjects at each location in the image. This means we learn different model parameters for many image locations involving methematical updates that involve many small matrix multiplications. In this project we want to investigate the potential for GPUs to help in the runtime of the algorithm.
| Objective | Approach and Plan | Progress and Next Steps |
|---|---|---|
|
|
|