DBP2:MIND:Roadmap
Back to NA-MIC Internal Collaborations, MIND DBP 2
Brain Lesion Analysis in Neuropsychiatric Systemic Lupus Erythematosus
Objective
This page describes the technology roadmap for lesion analysis in the NA-MIC Kit. Our objective is to create an end-to-end application allowing individual analysis of white matter lesions. This workflow applied to lupus patients is one of goals of the MIND DBP. The basic components necessary for this project are:
- Registration: co-registration of T1-weighted, T2-weighted, and FLAIR images
- Tissue segmentation: Should be multi-modality, correcting for intensity inhomogeneity and work on non-skull-stripped data.
- Lesion Localization: Each unique lesion should be detected and anatomical location summarized
- Lesion Load Measurement: Measure volume of each lesion, summarize lesion load by regions
- Time Series Analysis of white matter lesions: Compare lesions between multiple scans taken over time.
- Multi-scale Analysis of white matter lesions: Combine DTI, anatomical, and functional scans when analyzing lupus subjects.
- Tutorial: Documentation will be written for a tutorial and sample data sets will be provided
Roadmap
We obtained gray matter, white matter, CSF, and lesion maps for each subject based on T1-weighted, T2-weighted, and FLAIR images. Ultimately, the NA-MIC Kit will provide a workflow for individual and group analysis of lesions. It will be implemented as a set of Slicer3 modules that can be used interactively within the Slicer3 application as well as in batch on a computing cluster using BatchMake.
The current status of the main modules to be used are:
Registration
- Slicer has multiple image registration tools.
Lesion segmentation
- A Slicer module has been created that implements a novel lesion segmentation method based on multi-level morphometric feature classification. (Scully)
Manual tracing will serve as a bronze-standard:
- Slicer3 Manual tracing by clinically trained rater (Gasparovic/Bockholt)
Lesion Localization
- Slicer3 has tools for labeling white matter lesions and summarizing their anatomical location
Lesion Load Measurement
- Slicer3 has tools for measurement of labelled lesions
Performance characterization and validation
- Data will be collected at both 1.5 and 3T. Data at 1.5T will be obtained with the protocol utilized for the current project on lupus at UNM.
- Data at 3T will be obtained with sequences optimized for segmentation by the group at Utah.
- Comparisons will be based on the approach developed by Martin-Fernandez et al.
Tutorial
- 10 Externally sharable T1,T2,Flair,Lesion Map data-sets (NIFTI format) will be made available to the scientific community
- 5 subjects with lupus and 5 healthy normal volunteers
- A tutorial will be created that will guide end-users through each step needed to complete a lesion analysis in the NA-MIC kit
- A step-by-step guide will be produced for one example lupus case within the tutorial and one example healthy normal volunteer
- Measurements for all 10 subjects will also be provided so that users may compare their results with those of the tutorial
- 10 Externally sharable T1,T2,Flair,Lesion Map data-sets (NIFTI format) will be made available to the scientific community
Schedule
- Sequence Optimization and Data Collection
- 10/15/2007 T1, T2, FLAIR optimized sequences for Siemens 3T Trio Tim scanner (Jeremy, Bruce, Chuck)
- We will work with Bruce Fischl on using MEMPR, Mugler, and FLAIR sequences that have been optimized for maximum constrast and minimal geometric distortion across sequences
- DONE
- 3/31/2008 collection of 5 lupus subjects on clinical sequence and optimized 3T sequence (Chuck)
- collected 2 patients and 3 controls so far
- as of 3/10/2008 there are 3-4 additional patients scheduled, since january 1, we have had 4 patients not able to complete the protocol so far
- collected 2 patients and 3 controls so far
- 10/15/2007 T1, T2, FLAIR optimized sequences for Siemens 3T Trio Tim scanner (Jeremy, Bruce, Chuck)
- Co-Registration and Atlas Registration
- 1/31/2008 optimized mutual information registration for clinical sequences and optimized 3T sequences (Jeremy)
- we have tried the registration tools built-in to Slicer 3 EM-segment module so far
- DONE Brad has implemented this successfully into the EM-segment module
- 1/31/2008 optimized mutual information registration for clinical sequences and optimized 3T sequences (Jeremy)
- Lesion segmentation
- 3/17/2008 complete lesion segmentation methods for Slicer3 EM-segment, ITK itkEMS, ITK lesion segmentation, and Slicer3 manual tracing (Jeremy, Chuck, Vince Magnotta, Kilian/Brad, Marcel)
- Slicer3 EM-segment Module
- (Resolved) Currently has a bug that does not permit 3 channel segmentation Mantis ID#0000205
- (Resolved) Only using 2 channels such as T1/T2 or T1/FLAIR does not produce sensible/usable results
- (Resolved) Does not properly allow user to edit or change weighting of channels, this is crucial for hierarchical lesion classification. Mantis ID#0000204
- EMSegment has been run against collected lupus cases. Segmentations have improved but are still unusable as of 08/14/2008.
- Currently working with Killian on optimizing parameters in EMSegment to improve the lesion segmentation.
- Testing parameters and resulting segmentation on collected lupus cases.
- (Resolved) Currently has a bug that does not permit 3 channel segmentation Mantis ID#0000205
- ITK itkEMS stand-alone package
- Marcel and Mark are in active collaboration with getting this method to work on-site, we have encountered compilation and memory problems with the package on mac and linux environments so far
- 3/17/2008 update, marcel built a 64-bit linux version that appears to run successfully, current results are are shown here
- ITK lesion classification stand-alone package
- Vince Magnotta has develop an ITK-based 2 stage lesion segmentation method
- stage one uses k-means tissue classification into grey, white, or csf of co-registered T1/T2
- stage two use bayesian classifier into normal tissue or lesion using results of stage one and co-reigstered Flair
- current results of this method as of 3/17/2008 are shown here
- This method has been run against all cases as of 05/01/08. Currently working with Vince Magnotta to improve the performance.
- As of 08/14/2008, currently running a modified version of Vince's classifier that has been improved by replacing intensity thresholding with joint histogram feature vector occurrence thresholding.
- Vince Magnotta has develop an ITK-based 2 stage lesion segmentation method
- ITK K Nearest Neighbor Classifier
- Mark Scully is currently implementing a K-NN lesion classifier using ITK.
- This approach allows for on the fly additions to the classification model.
- The same segmentation approach is applicable to multiple lesion types.
- The resulting label map contains the percent likelihood that a given voxel is a lesion. This allows the clinician to adjust the thresholding to match their preference.
- On hold as of 1/12/2008 due to incomplete ITK algorithmic support
- Mark Scully is currently implementing a K-NN lesion classifier using ITK.
- ITK Custom Classifier
- Mark Scully is currently implementing a combined approach classifier using ITK.
- This approach allows for on the fly additions to the classification model.
- The same segmentation approach is applicable to multiple lesion types.
- This approach takes advantage of multiple approaches and advancements in the literature, including feature generation, Gentle Boosting based feature reduction, Artificial Immune System based filtering, and Bayesian classification.
- As of 1/01/2009 this method is functional and built into the lesion segmentation slicer3 applications.
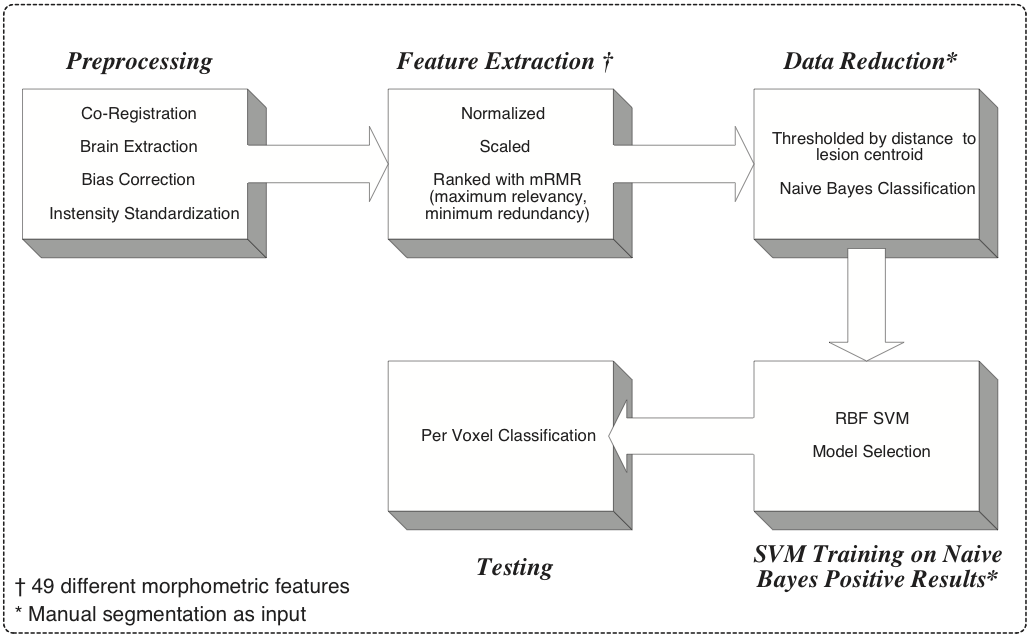
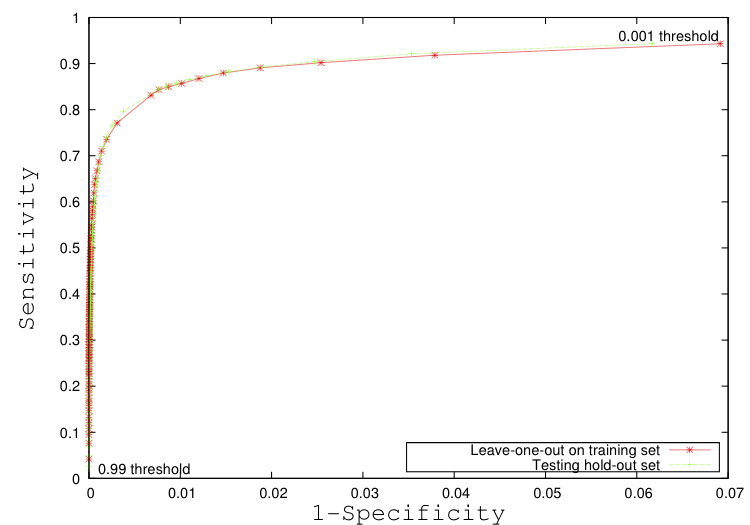
- Mark Scully is currently implementing a combined approach classifier using ITK.
- Slicer3 EM-segment Module
- 3/17/2008 complete lesion segmentation methods for Slicer3 EM-segment, ITK itkEMS, ITK lesion segmentation, and Slicer3 manual tracing (Jeremy, Chuck, Vince Magnotta, Kilian/Brad, Marcel)
- Slicer3 Manual Tracing
- Gasparovic has manually traced all tutorial cases with lesions using T2/Flair images following tracing guidelines and a qualitative neuroradiological review report
- 4/1/2008 create manual tracing guidelines documentation and share here
- Gasparovic has manually traced all tutorial cases with lesions using T2/Flair images following tracing guidelines and a qualitative neuroradiological review report
- Slicer3 Manual Tracing
- Lesion Localization
- 4/15/2008 complete lesion localizations and maps for Slicer3 EM-segment, ITK itkEMS, ITK lesion segmentation, and Slicer3 manual tracing (Jeremy, Chuck, Vince Magnotta, Steve)
- Lesion Measurement
- 4/20/2008 complete lesion measurements and regional lesion load summaries for Slicer3 EM-segment, ITK itkEMS, ITK lesion segmentation, and Slicer3 manual tracing (Jeremy, Chuck, Vince Magnotta, Steve)
- Performance characterization and validation
- 1/6/2008 report to 2008 NA-MIC AHM on progress for performance and validation of lesion segmentation methods for EM-segment, BRAINS2, manual, UNC packages (Jeremy)
- DONE
- 5/15/2008 submit abstract on reliability and summary of novel NA-MIC kit lesion analysis method (Jeremy, Chuck, Mark Scully, Brad, Kilian, others)
- plan is to submit SFN abstract and manuscript closely after
- 5/20/2008 analyze NIH study clinical sample using NA-MIC kit lesion analysis method (Jeremy, Chuck, Mark Scully)
- 7/1/2008 submit manuscript on clinical application of NA-MIC kit lesion analysis method (Jeremy, Chuck, Mark Scully, Carlos Roldan, Bill Sibbitt)
- 8/14/2008 submit manuscript on the lesion segmentation methods as applied to MS lesions for MICCAI 2008. (Mark, Jeremy, Chuck, Vince Magnotta)
- 1/11/2009 submit abstract on the new lesion segmentation methods as applied to Lupus lesions for HBM 2009.
- 3/08/2009 submit manuscript on the new lesion segmentation methods as applied to Lupus lesions for MICCAI 2009.
- 1/6/2008 report to 2008 NA-MIC AHM on progress for performance and validation of lesion segmentation methods for EM-segment, BRAINS2, manual, UNC packages (Jeremy)
- Incorporation of approach into NA-MIC kit
- 1/6/2008 Slicer3 Lesion Analysis Module that handles co-registration of T1, T2, and Flair (Mark Scully, Steve)
- DONE Brad completed this
- 5/1/2008 extend Slicer3 Lesion Analysis Module to handle lesion localization and measurement ('Mark Scully, Steve)
- 08/01/2008 currently locates all independent lesion clusters and identifies the anatomical regions, provided by a registered atlas, that the lesions overlap.
- 6/1/2008 extend Slicer3 Lesion Analysis Module to implement the ITK-based lesion analysis algorithm/method (EM-segment, ITK itkEMS, ITK lesion segmentation) with the best performance (Mark Scully, Steve)
- a demonstration will be given at the 2008 NA-MIC AHM--we would consider this version a prototype and would be testing, refining the module through the clinical application phase (Jeremy, Mark Scully)
- 6/15/2008 functioning prototype version of lesion analysis Slicer3 Module complete (Mark Scully, Steve)
- 8/14/2008 in the process of creating a module that "chains" together all of the steps in the lesion segmentation analysis pipeline.
- 1/03/2009 Newly developed lesion segmentation method with significantly increased performance incorporated into Slicer3 module.
- 1/6/2008 Slicer3 Lesion Analysis Module that handles co-registration of T1, T2, and Flair (Mark Scully, Steve)
- Tutorial and Data-sharing
- 6/20/2008 present tutorial to 2008 NA-MIC programming week (Mark Scully, Jeremy, Sonja)
- A logical project for programming week for Jeremy and Sonja to finish public version of tutorial
- 7/1/2008 make data sets and tutorial available to the scientific community (Mark Scully, Jeremy, Sonja)
- If we submit SFN abstract, we can advertise the availability during SFN Conference in November 2008
- We could think about advertising the roadmaps at the BIRN booth at SFN this year?
- If we submit SFN abstract, we can advertise the availability during SFN Conference in November 2008
- 10/15/2008 In the process of making the tutorial data available using XNAT.
- 6/20/2008 present tutorial to 2008 NA-MIC programming week (Mark Scully, Jeremy, Sonja)
Compare view of baseline and followup with color-coded lesion differences: http://www.na-mic.org/Wiki/index.php/File:CompareViewFlairLesionDiffWholeBrain.png
Diffusion tracts intersecting a lesion volume: http://www.na-mic.org/Wiki/index.php/File:LesionTractsNear.png
Axial slice with % predicted chance, thresholded predicted % chance, and manual tracing: http://www.na-mic.org/Wiki/index.php/File:Scully_Figure3.jpg
Publications
- Scully M, Anderson B, Lane T, Gasparovic C, Magnotta V, Sibbitt W, Roldan C, Kikinis R and Bockholt HJ (2010) An automated method for segmenting white matter lesions through multi-level morphometric feature classification with application to lupus. Front. Hum. Neurosci. 4:27. doi:10.3389/fnhum.2010.00027
- H. J. Bockholt, V. A. Magnotta, M. Scully, C. Gasparovic, B. Davis, K. Pohl, R. Whitaker, S. Pieper, C. Roldan, R. Jung, R. Hayek, W. Sibbitt, J. Sharrar, P. Pellegrino, R. Kikinis. A novel automated method for classification of white matter lesions in systemic lupus erythematosus. Presented at the 38th annual meeting of the Society for Neuroscience, Washingto, DC, 15 – 19 November 2008
- Scully M., Magnotta V., Gasparovic C., Pelligrimo P., Feis D., Bockholt H.J. 3D Segmentation In The Clinic: A Grand Challenge II at MICCAI 2008 - MS Lesion Segmentation. IJ - 2008 MICCAI Workshop - MS Lesion Segmentation. Available http://grand-challenge2008.bigr.nl/proceedings/pdfs/msls08/282_Scully.pdf
Team and Institute
- Co-PI: H Jeremy Bockholt (jbockholt at mrn.org)
- Co-PI: Charles Gasparovic (chuck at unm.edu)
- Software Engineer: Mark Scully (mscully at mrn.org)
- NA-MIC Engineering Contact: Steve Pieper, Isomics
- NA-MIC Algorithms Contact: Ross Whitaker, Utah
- Host Institues: The Mind Research Network and The University of New Mexico