Difference between revisions of "DBP2:UNC:Local Cortical Thickness Pipeline"
From NAMIC Wiki
| Line 6: | Line 6: | ||
We would like to create an end-to-end application within Slicer3 allowing individual and group analysis of local cortical thickness. | We would like to create an end-to-end application within Slicer3 allowing individual and group analysis of local cortical thickness. | ||
| − | == | + | <center> |
| + | {| | ||
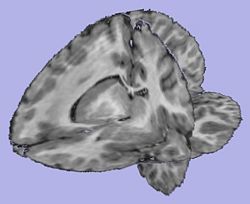
| + | |[[Image:T1Image.jpg|thumb|250px|T1-weighted skull-stripped image]] | ||
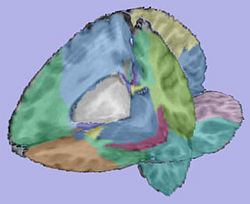
| + | |valign="center"|[[Image:Parcellation.jpg|thumb|250px|Parcellation image]] | ||
| + | |valign="center"|[[Image:WMThickness.jpg|thumb|250px|Cortical thickness on WM surface]] | ||
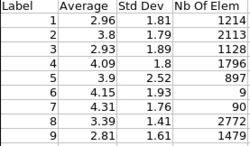
| + | |valign="center"|[[Image:ThicknessInformation.jpg|thumb|250px|Cortical thickness information]] | ||
| + | |} | ||
| + | </center> | ||
| − | + | == Pipeline overview == | |
Input: RAW images (T1-weighted, T2-weighted, PD-weighted images) | Input: RAW images (T1-weighted, T2-weighted, PD-weighted images) | ||
:* '''1. Tissue segmentation''' | :* '''1. Tissue segmentation''' | ||
| + | :** Probabilistic atlas-based automatic tissue segmentation via an Expectation-Maximization scheme | ||
:** Tool: itkEMS (UNC Slicer3 external module) | :** Tool: itkEMS (UNC Slicer3 external module) | ||
| − | :* '''2. Atlas-based ROI segmentation:''' subcortical structures, lateral ventricles, | + | :* '''2. Atlas-based ROI segmentation:''' subcortical structures, lateral ventricles, parcellation |
| − | :** 2.1 | + | :** 2.1 T1-weighted atlas deformable registration |
| − | :***Tool: RegisterImages (Slicer3 module) | + | :*** B-spline pipeline registration |
| + | :*** Tool: RegisterImages (Slicer3 module) | ||
:** 2.2. Applying transformations to the structures | :** 2.2. Applying transformations to the structures | ||
:*** Tool: ResampleVolume2 (Slicer3 module) | :*** Tool: ResampleVolume2 (Slicer3 module) | ||
| − | :* '''3. White matter map and | + | :* '''3. White matter map creation''' |
| + | :** Brainstem and cerebellum extraction | ||
| + | :** Adding subcortical structures except amygdala and hippocampus | ||
| + | :** Tool: ImageMath (NTIRC module) | ||
| + | :* '''4. White matter map post-processing''' | ||
| + | :** Largest component computation | ||
| + | :** White matter filling | ||
| + | :** Smoothing: Level set smoothing or weighted average filter | ||
| + | :** Connectivity enforcement (6-connectivity) | ||
| + | :** Tool: SegPostProcessB (Slicer3 external module) | ||
| + | :* '''5. Genus zero white matter map image and surface creation''' | ||
| + | :** Tool: GenusZeroImageFilter (UNC Slicer3 external module) | ||
| + | :* '''6. Gray matter map creation''' | ||
| + | :** Adding genus zero white matter map to gray matter segmentation (without cerebellum and brainstem) | ||
:** Tool: ImageMath | :** Tool: ImageMath | ||
| − | :* ''' | + | :* '''7. White matter surface inflation''' |
| − | :** Tool: | + | :** Tool: MeshInflation (UNC Slicer3 external module) |
| − | :* ''' | + | :* '''8. Cortical correspondence''' |
| + | :** Correspondence on inflated surface using particle system | ||
| + | :** Tool: ParticleCorrespondence (UNC Slicer3 external module) | ||
| + | :* '''9. Label map creation''' | ||
| + | :** Label map creation for cortical thickness computation (WM + GM + "CSF") | ||
:** Tool: ImageMath | :** Tool: ImageMath | ||
| − | :* ''' | + | :* '''10. Cortical thickness''' |
| − | :** | + | :** Asymmetric local cortical thickness or Laplacian cortical thickness |
| − | + | :** Tool: UNCCortThick or measureThicknessFilter (UNC Slicer3 external modules) | |
| − | :** Tool: | + | :* '''11. Group statistical analysis''' |
| − | :* ''' | + | :** Tool: QDEC Slicer module or StatNonParamPDM |
| − | :** Tool: | + | |
| − | + | == Download == | |
| − | |||
| − | === Usage | + | === Usage === |
| − | + | === Command line execution === | |
| + | === Step by step command line execution === | ||
| − | == | + | == Pipeline validation == |
| − | + | == Planning == | |
=== In progress === | === In progress === | ||
:* Cortical surface inflation: module in progress | :* Cortical surface inflation: module in progress | ||
| − | : | + | :* Iterative smoothing using relaxation operator (considering average vertex) and L2 norm of the mean curvature as a stopping criterion |
=== Future work === | === Future work === | ||
Revision as of 00:28, 14 December 2008
Home < DBP2:UNC:Local Cortical Thickness PipelineBack to UNC Cortical Thickness Roadmap
Objective
We would like to create an end-to-end application within Slicer3 allowing individual and group analysis of local cortical thickness.
Pipeline overview
Input: RAW images (T1-weighted, T2-weighted, PD-weighted images)
- 1. Tissue segmentation
- Probabilistic atlas-based automatic tissue segmentation via an Expectation-Maximization scheme
- Tool: itkEMS (UNC Slicer3 external module)
- 2. Atlas-based ROI segmentation: subcortical structures, lateral ventricles, parcellation
- 2.1 T1-weighted atlas deformable registration
- B-spline pipeline registration
- Tool: RegisterImages (Slicer3 module)
- 2.2. Applying transformations to the structures
- Tool: ResampleVolume2 (Slicer3 module)
- 2.1 T1-weighted atlas deformable registration
- 3. White matter map creation
- Brainstem and cerebellum extraction
- Adding subcortical structures except amygdala and hippocampus
- Tool: ImageMath (NTIRC module)
- 4. White matter map post-processing
- Largest component computation
- White matter filling
- Smoothing: Level set smoothing or weighted average filter
- Connectivity enforcement (6-connectivity)
- Tool: SegPostProcessB (Slicer3 external module)
- 5. Genus zero white matter map image and surface creation
- Tool: GenusZeroImageFilter (UNC Slicer3 external module)
- 6. Gray matter map creation
- Adding genus zero white matter map to gray matter segmentation (without cerebellum and brainstem)
- Tool: ImageMath
- 7. White matter surface inflation
- Tool: MeshInflation (UNC Slicer3 external module)
- 8. Cortical correspondence
- Correspondence on inflated surface using particle system
- Tool: ParticleCorrespondence (UNC Slicer3 external module)
- 9. Label map creation
- Label map creation for cortical thickness computation (WM + GM + "CSF")
- Tool: ImageMath
- 10. Cortical thickness
- Asymmetric local cortical thickness or Laplacian cortical thickness
- Tool: UNCCortThick or measureThicknessFilter (UNC Slicer3 external modules)
- 11. Group statistical analysis
- Tool: QDEC Slicer module or StatNonParamPDM
- 1. Tissue segmentation
Download
Usage
Command line execution
Step by step command line execution
Pipeline validation
Planning
In progress
- Cortical surface inflation: module in progress
- Iterative smoothing using relaxation operator (considering average vertex) and L2 norm of the mean curvature as a stopping criterion
Future work
- UNC Slicer3 external modules available on NITRIC
- Workflow for individual analysis (Slicer3 external module using BatchMake)
- Workflow for group analysis