DBP2:UNC:Regional Cortical Thickness Pipeline
Back to UNC Cortical Thickness Roadmap
Objective
We would like to create an end-to-end application within Slicer3 allowing individual and group analysis of regional cortical thickness.
General information
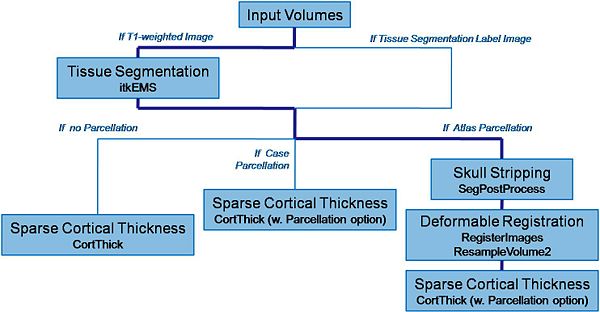
Pipeline description (steps)
Input: T1-weighted image, T2-weighted image, PD-weighted image
- 1. Tissue segmentation
- Tool: itkEMS (UNC Slicer3 external module)
- 2. Skull stripping
- Tool: SegPostProcess (UNC Slicer3 external module)
- 3. Deformable registration of pediatric regional atlas
- 3.1 Deformable registration of T1-weighted pediatric atlas
- Tool: RegisterImages (Slicer3 module)
- 3.2. Applying transformation to the parcellation map
- Tool: ResampleVolume2 (Slicer3 module)
- 3.1 Deformable registration of T1-weighted pediatric atlas
- 4. Cortical Thickness
- Tool: CortThick (UNC Slicer3 module)
- 1. Tissue segmentation
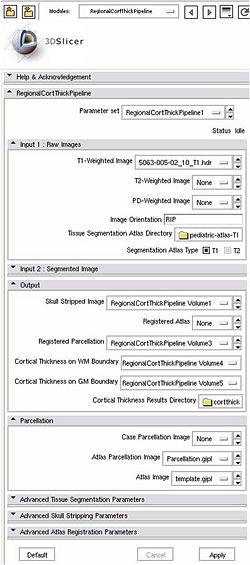
All the tools used in the current pipeline are Slicer3 modules, some of them being UNC external modules. The user can thus compute an individual regional cortical thickness analysis by running the 'RegionalCortThickPipeline' module, either within Slicer3 or as a command line.
Usage (Command Line)
Inputs: T1-weighted image, T1-weigthed atlas, regional atlas (parcellation map)
A. Pipeline command line
RegionalCortThickPipeline --T1 Image_T1.gipl --segAtlasDir TissueSegmentationAtlasDirectory/ --atlas Atlas.gipl --atlasParcellation Parcellation.gipl --SaveWM WMCorticalThicknessMap --SaveGM GMCorticalThicknessMap
B. Step by step command line
- 1. Tissue segmentation
- Input: EMSparam.xml
- Output: Image_Corrected_EMS.gipl, Label.gipl
- 1. Tissue segmentation
itkEMSCLP --XMLFile EMSparam.xml
- 2. Skull stripping
- Input: Label.gipl, Image_Corrected_EMS.gipl
- Output: Image_Corrected_EMS_Stripped.gipl, BinaryMask.gipl (optional)
- 2. Skull stripping
SegPostProcessCLP Label.gipl Image_Corrected_EMS_Stripped.gipl --skullstripping Image_Corrected_EMS.gipl
- 3. Deformable registration of pediatric regional atlas
- 3.1 Deformable registration of T1-weighted pediatric atlas
- Input: Atlas.gipl, Image_Corrected_EMS_Stripped.gipl
- Output: Atlas_Registered.gipl, Atlas_Registered_Transform.txt
- 3.1 Deformable registration of T1-weighted pediatric atlas
- 3. Deformable registration of pediatric regional atlas
RegisterImages Image_Corrected_EMS_Stripped.gipl Atlas.gipl –resampledImage Atlas_Registered.gipl –saveTransform Atlas_Registered_Transform.txt –registration PipelineBSpline
- 3.2. Applying transformation to the parcellation map
- Input: Parcellation.gipl, Atlas_Registered_Transform.txt, Image_Corrected_EMS_Stripped.gipl
- Output: Parcellation_Registered.gipl
- 3.2. Applying transformation to the parcellation map
ResampleVolume2 Parcellation.gipl Parcellation_Registered.gipl -f Atlas_Registered_Transform.txt -i nn -R Image_Corrected_EMS_Stripped.gipl
- 4. Cortical Thickness
- Input: Parcellation_Registered.gipl, Label.gipl
- Output: CortThick_Result_Dir/, WMCorticalThicknessMap, GMCorticalThicknessMap
- 4. Cortical Thickness
CortThickCLP CortThick_Result_Dir/ --par Parcellation_Registered.gipl --inputSeg Label.gipl --SaveWM WMCorticalThicknessMap --SaveGM GMCorticalThicknessMap
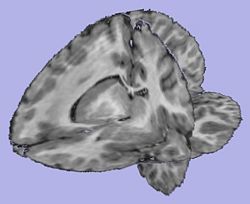
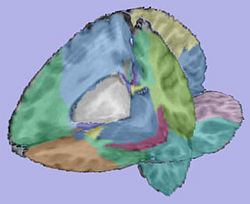
Analysis on a small pediatric dataset
Tests have been computed on a small pediatric dataset, including 2 years old and 4 years old cases
- 2 Autistic cases
- 1 developmental delay
- 1 normal control
In progress
- Workflow for individual analysis (Slicer3 external module using BatchMake)
Future work
- UNC Slicer3 external modules available on NITRIC
- Pediatric atlas (T1-weighted image and parcellation map) available to the community (XNAT?)
- Workflow for group analysis