NA-MIC External Collaborations
This section describes external collaborations with NA-MIC that are funded by NIH under the "Collaboration with NCBC" PAR. (Details for this funding mechanism are provided here).
Contents
- 1 PAR-05-063: R01CA124377 An Integrated System for Image-Guided Radiofrequency Ablation of Liver Tumors
- 2 PAR-05-063: R01EB005973 Automated FE Mesh Development
- 3 PAR-05-063: R01AA016748 Measuring Alcohol and Stress Interaction with Structural and Perfusion MRI
- 4 PAR-05-057: BRAINS Morphology and Image Analysis
- 5 Children's Pediatric Cardiology Collaboration with SCI/SPL/Northeastern
PAR-05-063: R01CA124377 An Integrated System for Image-Guided Radiofrequency Ablation of Liver Tumors
This project is a NCBC collaboration grant. Collaborators include Kevin Cleary and Noby Hata. The master page for this collaboration is here, while the rest of this section contains links to the specific active projects in this collaboration.
Projects
To be added.
PAR-05-063: R01EB005973 Automated FE Mesh Development
This project is a NCBC collaboration grant. Collaborators include Nicole Grosland, Vincent Magnotta, Steve Pieper, and Simon Warfield. The master page for this collaboration is here, while the rest of this section contains links to the specific active projects in this collaboration.
Meshing Algorithms
- Voxel Meshing Module (Iowa)
- Novel Hexahedral Meshing Algorithms (Iowa)
- Hex vs Tet Mesh Comparisons (Iowa/Isomics/BWH)
Automated Segmentation
Image Registration
- Evaluation_of_Inter-Modality_Registration (Iowa/Isomics)
- Inter-slice Motion Correction for fMRI (OSU)
Validation
Mesh Quality Visualization
- FE Mesh Quality Visualization (Iowa/Isomics)
- Standalone FE Mesh Quality Viewer
- Mesh Quality Command Line Module
PAR-05-063: R01AA016748 Measuring Alcohol and Stress Interaction with Structural and Perfusion MRI
This project is a NCBC collaboration grant. Collaborators include James B Daunais, Robert Kraft, Chris Wyatt, William Wells, and Kilian Pohl. The master page for this collaboration is here, while the rest of this section contains links to the specific active projects in this collaboration.
Rhesus Probabalistic Atlas
Segmentation
PAR-05-057: BRAINS Morphology and Image Analysis
This project is not an NCBC collaboration grant, but instead a Continued Development and Maintenance of Software grant. The intent of this application is to update the BRAINS image analysis software developed at the University of Iowa. The collaborators include Vincent Magnotta, Hans Johnson, Jeremy Bockholt, and Nancy Andreasen. There are three main thrusts of this application as outlined below.
Implement a Automated Brain Analysis Pipeline
Code Refactoring for Cross Platform Support
- Refactor Current Modules using ITK and VTK
- Build GUI using Slicer3 Interface
- Use Python as Scripting Language
Validation of Pipeline Using MIND MCIC Sample
- Evaluate MIND MCIC Human Phantom Sample Using Automated Pipeline
- Evaluate Differences between Patients with Schizophrenia and Normal Controls using Automated Pipeline
Children's Pediatric Cardiology Collaboration with SCI/SPL/Northeastern
|
|
Introduction
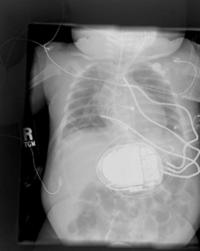
Placement of Implantable Cardiac Defibrillators(ICDs)is a unique and challenging problem for children due to the variety of shapes and sizes, ranging from neonate to adolescent. As a result, a variety of novel implant techniques have been employed. Although these have generally been successful inasmuch as they result in a clinically acceptable defibrillation threshold, nothing is known about the mechanisms by which this threshold is attained, the optimal geometries for defibrillation, and whether unsafe electric field strengths are a result of novel implant approaches. Finite element modeling has been shown in adult torso models to correlate well with clinical results. Our goal is to model defibrillation in child torso models to gain insight into this important problem. We are also interested in developing new orientations in adults with the goal of lowering DFTs and providing options in patients with contraindications to standard techniques.
Goals of Project
1. Create 3D models of children based on CT and MRI datasets for modeling internal and external defibrillation in the SCIRun environment. The processes required address the larger question of how to take any CT or MRI DICOM dataset, segment it into various label maps, combine those label maps in a hierarchical manner, then import and utilize them in the SCIRun/BioPSE environment. This represents part of an expanding collaboration between SCI and SPL to integrate open source tools to allow creation, visualization, and computational modeling of image based 3D models.
2. Create modules which allow insertion of electrode shapes into finite element models in in the SCIRun environment.
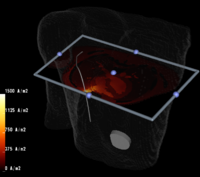
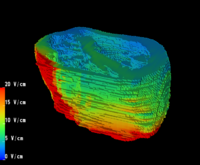
3. Utilize the above innovations to model the placement of internal defibrillator electrodes to maximize efficacy, minimize potential cardiac damage, and gain further insight into optimizing defibrillation in children of various sizes as shown above.
Slicer and SPL Tools in the Project
Outline of Torso Segmentation Problem
Segmentation Tools Currently Utilized
Tools In Development and Wishlist