Projects:ConnectivityAtlas
Back to NA-MIC Collaborations, MIT Algorithms,
Atlas building
this and that
Description
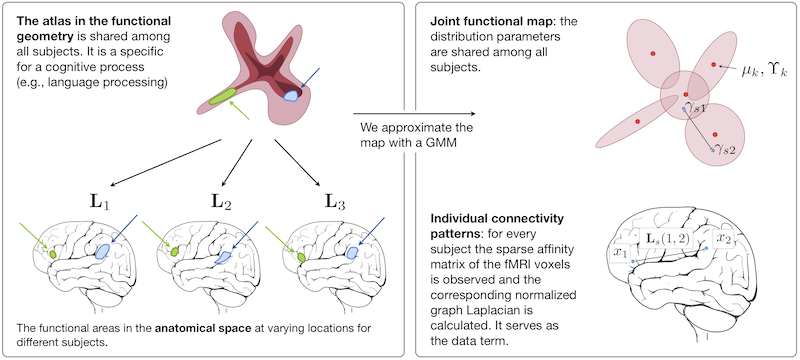
We construct an atlas that captures functional connectivity characteristics of a cognitive process from a population of individuals. The functional connectivity is encoded in a low-dimensional embedding space derived from a diffusion process on a graph that represents correlations of fMRI time courses. The atlas is represented by a common prior distribution for the embedded fMRI signals of all subjects. The atlas is not directly coupled to the anatomical space, and can represent functional networks that are variable in their spatial distribution. We derive an algorithm for fitting this generative model to the observed data in a population. Our results in a language fMRI study demonstrate that the method identifies coherent and functionally equivalent regions across subjects.
Experiments
Conclusion
Literature
[1] X. Artaechevarria, A. Munoz-Barrutia, and C. Ortiz de Solorzano. Combination strategies in multi-atlas image segmentation: Application to brain MR data. IEEE Tran. Med. Imaging, 28(8):1266 – 1277, 2009.
[2] R.A. Heckemann, J.V. Hajnal, P. Aljabar, D. Rueckert, and A. Hammers. Automatic anatomical brain MRI segmentation combining label propagation and decision fusion. Neuroimage, 33(1):115–126, 2006.
[3] T. Rohlfing, R. Brandt, R. Menzel, and C.R. Maurer. Evaluation of atlas selection strategies for atlas-based image segmentation with application to confocal microscopy images of bee brains. NeuroImage, 21(4):1428–1442, 2004.
[4] Freesurfer Wiki. http://surfer.nmr.mgh.harvard.edu.
Key Investigators
- MIT: Georg Langs, Andy Sweet, Danial Lashkari, Polina Golland
- Harvard: Yanmei Tie, Laura Rigolo, Alexandra Golby
Publications
NA-MIC Publications Database on Nonparametric Models for Supervised Image Segmentation