Difference between revisions of "Projects:MultiscaleShapeSegmentation"
| Line 1: | Line 1: | ||
| − | |||
| − | |||
Back to [[NA-MIC_Collaborations|NA-MIC_Collaborations]], [[Algorithm:GATech|Georgia Tech Algorithms]], [[Algorithm:UNC|UNC Algorithms]] | Back to [[NA-MIC_Collaborations|NA-MIC_Collaborations]], [[Algorithm:GATech|Georgia Tech Algorithms]], [[Algorithm:UNC|UNC Algorithms]] | ||
| − | + | = Multiscale Shape Segmentation = | |
To represent multiscale variations in a shape population in order to drive the segmentation of deep brain structures, such as the caudate nucleus or the hippocampus. | To represent multiscale variations in a shape population in order to drive the segmentation of deep brain structures, such as the caudate nucleus or the hippocampus. | ||
| − | '' | + | = Description = |
| + | |||
| + | ''Shape Representation and Prior'' | ||
| − | |||
The overview of our shape representation is given in Figure 1. Our technique defines a multiscale parametric model of surfaces belonging to the same population using a compact set of spherical wavelets targeted to that population (Figure 2). We further refine the shape representation by separating into groups wavelet coefficients that describe independent global and/or local biological variations in the population, using spectral graph partitioning. We then learn a prior probability distribution induced over each group to explicitly encode these variations at different scales and spatial locations (Figure 4) [1]. | The overview of our shape representation is given in Figure 1. Our technique defines a multiscale parametric model of surfaces belonging to the same population using a compact set of spherical wavelets targeted to that population (Figure 2). We further refine the shape representation by separating into groups wavelet coefficients that describe independent global and/or local biological variations in the population, using spectral graph partitioning. We then learn a prior probability distribution induced over each group to explicitly encode these variations at different scales and spatial locations (Figure 4) [1]. | ||
| Line 21: | Line 20: | ||
We applied our algorithm to the caudate nucleus, a brain structure of interest in the study of schizophrenia [2]. Our validation shows our algorithm is computationally efficient and outperforms the Active Shape Model (ASM) algorithm, by capturing finer shape details. | We applied our algorithm to the caudate nucleus, a brain structure of interest in the study of schizophrenia [2]. Our validation shows our algorithm is computationally efficient and outperforms the Active Shape Model (ASM) algorithm, by capturing finer shape details. | ||
| − | + | = Key Investigators = | |
| + | |||
| + | * Georgia Tech: Delphine Nain, Aaron Bobick, Allen Tannenbaum | ||
| + | * Harvard SPL: Steven Haker | ||
| + | |||
| + | |||
| + | = Publications = | ||
| + | |||
* Nain D, Haker S, Bobick A, Tannenbaum A. Multiscale 3D Shape Analysis using Spherical Wavelets. Proc MICCAI, Oct 26-29 2005, p 459-467. | * Nain D, Haker S, Bobick A, Tannenbaum A. Multiscale 3D Shape Analysis using Spherical Wavelets. Proc MICCAI, Oct 26-29 2005, p 459-467. | ||
* Nain D, Haker S, Bobick A, Tannenbaum A. Shape-driven 3D Segmentation using Spherical Wavelets. Proc MICCAI, Oct 2-5, 2006. | * Nain D, Haker S, Bobick A, Tannenbaum A. Shape-driven 3D Segmentation using Spherical Wavelets. Proc MICCAI, Oct 2-5, 2006. | ||
| − | |||
| − | |||
| − | |||
| − | |||
''Collaborators'' | ''Collaborators'' | ||
| Line 36: | Line 38: | ||
* Core 3: James Levitt, Marc Niethammer, Sylvain Bouix, Martha Shenton (Harvard PNL) | * Core 3: James Levitt, Marc Niethammer, Sylvain Bouix, Martha Shenton (Harvard PNL) | ||
| − | + | = Links = | |
* Paper presented in [[MICCAI_2006|MICCAI 2006, Copenhagen, October 2 - 4, 2006 ]] | * Paper presented in [[MICCAI_2006|MICCAI 2006, Copenhagen, October 2 - 4, 2006 ]] | ||
Revision as of 21:11, 21 September 2007
Home < Projects:MultiscaleShapeSegmentationBack to NA-MIC_Collaborations, Georgia Tech Algorithms, UNC Algorithms
Multiscale Shape Segmentation
To represent multiscale variations in a shape population in order to drive the segmentation of deep brain structures, such as the caudate nucleus or the hippocampus.
Description
Shape Representation and Prior
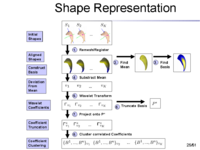
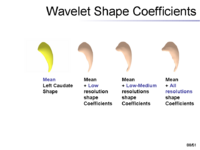
The overview of our shape representation is given in Figure 1. Our technique defines a multiscale parametric model of surfaces belonging to the same population using a compact set of spherical wavelets targeted to that population (Figure 2). We further refine the shape representation by separating into groups wavelet coefficients that describe independent global and/or local biological variations in the population, using spectral graph partitioning. We then learn a prior probability distribution induced over each group to explicitly encode these variations at different scales and spatial locations (Figure 4) [1].
Segmentation Based on this representation, we derive a parametric active surface evolution using the multiscale prior coefficients as parameters for our optimization procedure to naturally include the prior for segmentation. Additionally, the optimization method can be applied in a coarse-to-fine manner.
Results We applied our algorithm to the caudate nucleus, a brain structure of interest in the study of schizophrenia [2]. Our validation shows our algorithm is computationally efficient and outperforms the Active Shape Model (ASM) algorithm, by capturing finer shape details.
Key Investigators
- Georgia Tech: Delphine Nain, Aaron Bobick, Allen Tannenbaum
- Harvard SPL: Steven Haker
Publications
- Nain D, Haker S, Bobick A, Tannenbaum A. Multiscale 3D Shape Analysis using Spherical Wavelets. Proc MICCAI, Oct 26-29 2005, p 459-467.
- Nain D, Haker S, Bobick A, Tannenbaum A. Shape-driven 3D Segmentation using Spherical Wavelets. Proc MICCAI, Oct 2-5, 2006.
Collaborators
- Core 1: Martin Styner (UNC)
- Core 2: Jim Miller (GE), Luis Ibanez (Kitware)
- Core 3: James Levitt, Marc Niethammer, Sylvain Bouix, Martha Shenton (Harvard PNL)