Projects:RegistrationLibrary:RegLib C10
From NAMIC Wiki
Home < Projects:RegistrationLibrary:RegLib C10
Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
Slicer Registration Library Exampe #10: Co-registration of probabilistic tissue atlas for subsequent EM segmentation
Objective / Background
This is an example of sparse atlas co-registration. Not all atlases have an associated reference image that can be used for registration. Because the atlas represents a map of a particular tissue class probability, its contrast differs significantly from the target image.
Keywords
MRI, brain, head, inter-subject, probabilistic atlas, atlas-based segmentation
Input Data
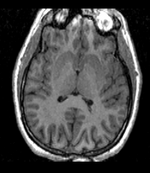
 reference/fixed : T1w axial, 1mm resolution in plane, 3mm slices
reference/fixed : T1w axial, 1mm resolution in plane, 3mm slices moving: Probabilistic Tissue atlas,
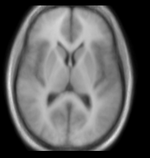
moving: Probabilistic Tissue atlas,
x 36 x 9
Methods
- build brain mask for fixed image using Skull Stripping module. Settings: 100 iterations, 20 subdivisions. New Volume: RegLib_C10_MRI_AtlasSegmentation_fixed_mask
- manually edit brain mask with Editor. required manual fix at frontal and occipital lobe
- run Register Images , Settings:
- Fixed Image:
- Moving Image:
- Resample Image:
- Load Transform:
- Save Transform: RegLib_C10_MRI_AtlasSegmentation_Xform_Affine_wmsk
- Initialization: Centers of Mass,
- Registration: PipelineAffine
- Expected offset: 10
- Expected Rotation: 0.2
- Expected Scale: 0.1
- Expected Skew: 0.05
- Fixed Image Mask: RegLib_C10_MRI_AtlasSegmentation_fixed_mask
- Affine Max Iteration: 80
- Affine Sampling Ratio: 0.05
- (alternatively automated Affine Registration: Register Images Multires (Slicer 3.5) also produces good results
- run '"Deformable B-spline Registration'" module. Settings:
- Grid Size: 5
- Histogram Bins: 50,
- Spatial Samples: 50000,
- initial transform: RegLib_C10_MRI_AtlasSegmentation_Xform_Affine_wmsk
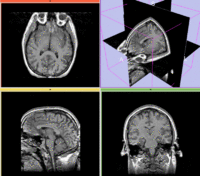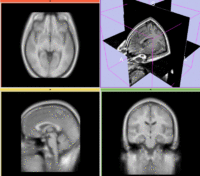
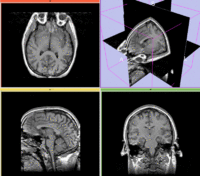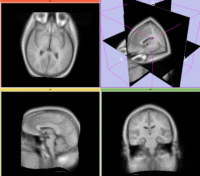
Registration Results
Download
- download entire package (Data,Tutorial, Solution, zip file xx MB)
- Presets
- Tutorial only
- Image Data only
Discussion: Registration Challenges
- Because the atlas represents a map of a particular tissue class probability, its contrast differs significantly from the target image.
- The two images may have strong differences in voxel sizes and voxel anisotropy. If the orientation of the highest resolution is not the same in both images, finding a good match can be difficult.
- The two images represent different anatomies, a non-rigid registration is required
Discussion: Key Strategies
- Because of the strong differences in image contrast, Mutual Information is recommended as the most robust metric.
- masking (skull stripping) is highly recommended to obtain good results.
- because speed is not that critical, we increase the sampling rate from the default 2% to 15%.
- we also expect larger differences in scale & distortion than with regular structural scans: so we significantly (2x-3x) increase the expected values for scale and skew from the defaults.
- a good affine alignment is important before proceeding to non-rigid alignment to further correct for distortions.