Projects:RegistrationLibrary:RegLib C31
From NAMIC Wiki
Home < Projects:RegistrationLibrary:RegLib C31
v3.6.1
Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
Contents
v3.6.1  Slicer Registration Library Case xx: TBI
Slicer Registration Library Case xx: TBI
Input
Modules
- Slicer 3.6.1 recommended modules: BrainsFit
Objective / Background
We seek to align two exams (acute baseline and follow-up) as well as all series within exams into a common space for direct evaluation of regional change.
Input Data
- inter-exam
- reference/fixed : T1_e1 baseline exam of acute TBI , 512 x 512 x 160 ; 0.5 x 0.5 x 1 mm voxel size, axial acquisition, RAS orientation.
- moving:T1_e2 follow-up exam of acute TBI , 512 x 512 x 160 ; 0.5 x 0.5 x 1 mm voxel size, axial acquisition, RAS orientation.
- intra-exam
Registration Challenges
- we have multiple nested transforms: each exam is co-registered within itself, and then the exams are aligned to eachother
- strong pathology (hemorrhage?) present at different amounts in the two exams
Key Strategies
- we first register the scans within each exam to the T1
- second we register the follow-up T1 scan to the baseline T1
- we then nest the first alignment within the second
Procedure
- Load & Center
- Load T1 MPRAGE datasets via Load Volume...
- volumes are note well centered, i.e. their physical origin defined in the image header differs; we therefore first center both images:
- Go to Volumes module, open Info tab
- From Active Volume menu, select each image in turn, then click the Center Image button
- note how Image origin changes for T1_e1 from 121,-97,-97 to 128,-128,-80 and for T1_e2 from 129,-106,-66 to 128,-128,-80
- now that both images have same center their initial misalignment can be seen by placing T1_e1 in the background and T1_e in the foreground and using the toggle-switch
- Align Exam 2 to Exam 1: T1 1st pass: unmasked
- Open Registration / BRAINSFit module
- Fixed Image: T1_e1, moving image: T1_e2
- Registration Phases: select "Initialize with CenterOfHeadAlign", Include Rigid, "IncludeScaleVersor3D" and Include Affine
- Output Settings: under SlicerLinear Transform, select "Create New Linear Transform, then select Rename" and rename it to Xf1_e2-e1_Affine
- Registration Parameters: increase the Number of Samples field to 200,000
- Leave all other settings at defaults & Click: Apply. Registration should complete within ~ 30 seconds
- Go back to the Data module: you should see the T1_e2 image moved under the newly created transform
- Select "T1_e1" as background and T2_e2 as new foreground, toggle to see alignment
- due to the strong deformations an affine alignment alone does not achieve a perfect match. We can extend the DOF to non-rigid BSpline transforms, but must be aware that doing so might remove differences of interest:
- Align Exam 2: 2nd pass: nonrigid
- Open Registration / BRAINSFit module
- Fixed Image: T1_e1, moving image: T1_e2
- From "Initialize with previously generated transform" & select the above affine Xf1_e2-e1_Affine
- Registration Phases: select only Include BSpline.
- Output Settings: under SlicerBSpline Transform, select "Create New BSpline Transform, then select Rename" and rename it to Xf2_e2-e1_BSpl
- Align Within Exam 1:
- Select Load Volume... and open the GRE, FLAIR and DWI images, respectively
- Open Registration / BRAINSFit module
- Select the presets Xf3... to Xf9... or use settings below:
- Registration phases: select Initialize with Center of Head Align, include rigid, include ScaleVersor3D and include Affine
- increase Number of Samples in Registration Parameters to 200,000
- Registration Parameters: increase the Number of Samples field to 200,000
- Leave all other settings at defaults & Click: Apply. Registration should complete within 3 minutes.
- You should see the earlier residual misalignment reduced, but strong distortions applied to the e2 image (see results below)
Registration Results
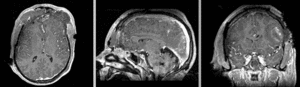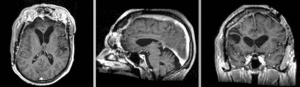
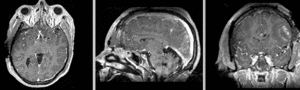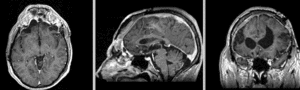 unregistered exam 1 & 2
unregistered exam 1 & 2
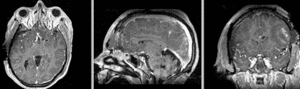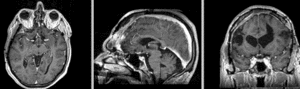 Exam 2 aligned to Exam 1 (affine only)
Exam 2 aligned to Exam 1 (affine only)
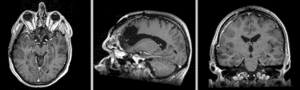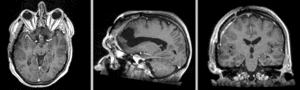
 Exam 2 aligned to Exam 1 (affine+BSpline)
Exam 2 aligned to Exam 1 (affine+BSpline)
 BSpline deformation only of Exam 2
BSpline deformation only of Exam 2