Difference between revisions of "Projects:RegistrationLibrary:RegLib C46"
From NAMIC Wiki
m (Text replacement - "http://www.slicer.org/slicerWiki/index.php/" to "https://www.slicer.org/wiki/") |
|||
| (23 intermediate revisions by one other user not shown) | |||
| Line 3: | Line 3: | ||
[[Projects:RegistrationDocumentation:UseCaseInventory|Back to Registration Use-case Inventory]] <br> | [[Projects:RegistrationDocumentation:UseCaseInventory|Back to Registration Use-case Inventory]] <br> | ||
| − | ==<small>v3.6. | + | ==<small>v3.6.3</small> [[Image:Slicer3-6Announcement-v1.png|150px]] Slicer Registration Library Case #46: 2D Cine MRI of Breathing Cycle == |
=== Input === | === Input === | ||
| Line 17: | Line 17: | ||
=== Modules === | === Modules === | ||
| − | *'''Slicer 3.6.1 recommended modules: [ | + | *'''Slicer 3.6.1 recommended modules: [https://www.slicer.org/wiki/Modules:BRAINSFit BrainsFit]''' |
===Objective / Background === | ===Objective / Background === | ||
| Line 25: | Line 25: | ||
===Input Data=== | ===Input Data=== | ||
| − | *reference/fixed : 2D coronal MRI 256x256 | + | *reference/fixed : 2D coronal MRI 256x256 , FIESTA sequence ,1.5 x 1.5 x 15mm voxel size |
| − | *moving: 2D coronal time series, 200 images total, each | + | *moving: 2D coronal time series, 200 images total, each as fixed above |
| − | === Overall Strategy === | + | === Notes / Overall Strategy === |
| + | *because the torso is already well registered and does not move across the sequence, a global affine/rigid registration is of no use. Instead we seek to assess the internal displacements caused across the breathing cycle. We therefore choose a low-DOF BSpline model. | ||
| + | *Because the implementation of the BRAINSfit module used here expects a 3D volume, we choose a slightly larger grid and make it isotropic, although the 3rd dimension of the grid is not used. We expect the optimizer in the registration to not move outside the slice plane and simply return 0 deformations there. | ||
| + | *An alternative strategy would be to register each image to its immediate previous timepoint, and then integrating the displacements. This has the advantage that nonrigid registration tends to be better for smaller differences, but it bears the risk of accumulating error and hence a lower robustness. | ||
| + | *A related option to improve precision would be to initialize each registration with the result of the previous one. Unfortunately the BRAINSfit registration modules do not (yet) allow to initialize with anything other than an affine. | ||
| + | *The additional information/constraint, that the displacements defined by the series of registrations should describe a smooth motion that is contiguous and differentiable, has not been exploited in this approach. This could be done a posteriori by applying a temporal filter across the transforms or fitting an appropriate model curve to each node displacement. | ||
| + | *Overall strategy: | ||
:#select one frame within the breathing cycle as reference frame | :#select one frame within the breathing cycle as reference frame | ||
:#compute non-rigid BSpline registration of all images to the reference frame | :#compute non-rigid BSpline registration of all images to the reference frame | ||
| Line 48: | Line 54: | ||
##click: Apply; (runtime < 10 sec. on MacPro) | ##click: Apply; (runtime < 10 sec. on MacPro) | ||
##adjust grid size until registration is acceptable | ##adjust grid size until registration is acceptable | ||
| + | ##you can see the commandline text of the registration performed by opening the ''Window/Error Log'' window and clicking on ''BRAINSfit commandline input'' | ||
*'''Phase II: Batch Run''' | *'''Phase II: Batch Run''' | ||
#open a terminal window | #open a terminal window | ||
| Line 58: | Line 65: | ||
## note that because input image is DICOM, and images are 2D only, each image of the time series must be in its own directory, otherwise Slicer will read them as a volume. | ## note that because input image is DICOM, and images are 2D only, each image of the time series must be in its own directory, otherwise Slicer will read them as a volume. | ||
##paste all commands into a terminal window, or copy into a shell script and execute. | ##paste all commands into a terminal window, or copy into a shell script and execute. | ||
| + | *'''Phase III: Evaluate Transform files''' | ||
##upon completion, read the transform files with an editor and extract the displacements of interest | ##upon completion, read the transform files with an editor and extract the displacements of interest | ||
| + | ##The ITK transform files describe displacements at the grid nodes, many of which are outside the region of interest. Because BRAINSfit pads the grid with 1 voxel, the grid returned is actually 8x8x8. We use only the plane (*,*,4) for analysis and discard the y-direction displacements, which as expected are all zero. | ||
| + | ##for details on the ITK transform format see the [https://www.slicer.org/wiki/Slicer-3-6-FAQ#What_is_the_Meaning_of_.27Fixed_Parameters.27_in_the_transform_file_.28.tfm.29_of_a_BSpline_registration_.3F '''FAQ''' here] | ||
| + | ##To obtain displacements at arbitrary coordinates, interpolate the transform into a deformation field, e.g. using this module (details [https://www.slicer.org/wiki/Slicer-3-6-FAQ#How_can_I_convert_a_BSpline_transform_into_a_deformation_field.3F FAQ here]): | ||
| + | /Applications/Slicer3.6.3/lib/Slicer3/Plugins/BSplineToDeformationField | ||
=== Registration Results=== | === Registration Results=== | ||
| + | {| style="color:#bbbbbb; " cellpadding="10" cellspacing="0" border="0" | ||
| + | |[[Image:RegLib_C46_Thumb2.gif|300px|lleft|moving input]] | ||
| + | |[[Image:RegLib_C46_Moving_animReg.gif|300px|lleft|moving input after registration (only frames 1-27 shown)]] | ||
| + | |[[Image:RegLib_C46_Displacements.png|300px|lleft|displacements]] | ||
| + | |- | ||
| + | |unregistered moving series | ||
| + | |moving input after registration (only frames 1-27 shown) | ||
| + | |[[Media:RegLib_C46_Displacements.pdf|displacements (PDF)]] | ||
| + | |} | ||
| + | |||
| − | |||
| − | |||
===Download === | ===Download === | ||
*Data | *Data | ||
| − | **[[Media:RegLib_C46_Data.zip |'''RegLib_C46_Data''' | + | **[[Media:RegLib_C46_Data.zip |'''RegLib_C46_Data''' : DICOM 2D series <small> (DICOM files, zip file 18 MB) </small>]]''' |
| − | + | *Results | |
| + | **[[Media:RegLib_C46_RegistrationTransforms.zip |'''RegLib_C46_RegistrationTransforms.zip''' : the 200 BSpline registration transforms <small> (ITK transform (text) files, zip file 274 kB) </small>]]''' | ||
| + | **[[Media:RegLib_C46_MovingResampled.zip |'''RegLib_C46_MovingResampled.zip''' : the 200 registered image stack <small> (NRRD image files, zip file 16MB) </small>]]''' | ||
=== Acknowledgments === | === Acknowledgments === | ||
| − | Thanks to Amir Owrangi and the [http://www.robarts.ca Robarts Institute] for sharing the registration problem and data. | + | Thanks to Amir Owrangi and [http://www.imaging.robarts.ca/~gep Dr.Grace Parraga] from the [http://www.robarts.ca Robarts Institute] for sharing the registration problem and data. |
Latest revision as of 18:07, 10 July 2017
Home < Projects:RegistrationLibrary:RegLib C46Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
Contents
v3.6.3  Slicer Registration Library Case #46: 2D Cine MRI of Breathing Cycle
Slicer Registration Library Case #46: 2D Cine MRI of Breathing Cycle
Input
|
|
|
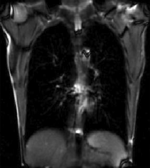
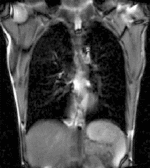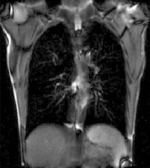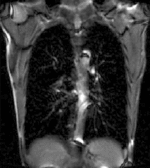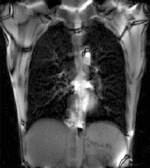
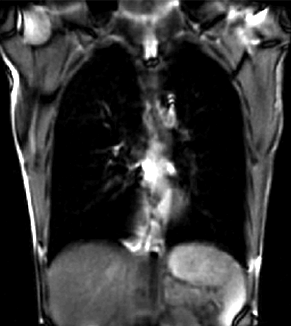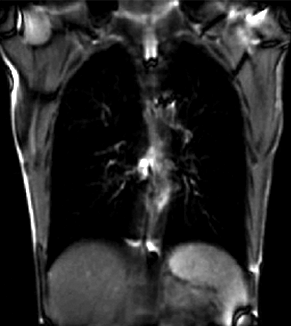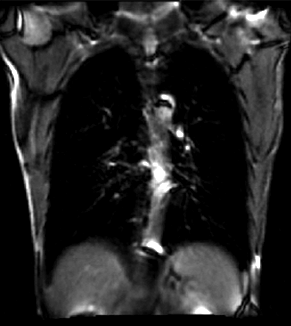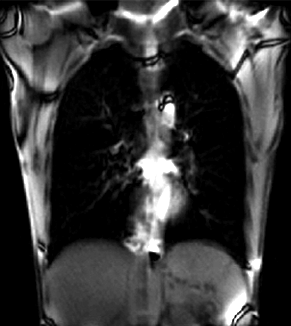
| fixed image/target | moving image |
Modules
- Slicer 3.6.1 recommended modules: BrainsFit
Objective / Background
We seek to study the motion of the lung during a breathing cycle by co-registering all of the images in a sequence with a reference, and then extract the motion from the registration displacement field.
Keywords
MRI, lung, 2D, time-series, deformation
Input Data
- reference/fixed : 2D coronal MRI 256x256 , FIESTA sequence ,1.5 x 1.5 x 15mm voxel size
- moving: 2D coronal time series, 200 images total, each as fixed above
Notes / Overall Strategy
- because the torso is already well registered and does not move across the sequence, a global affine/rigid registration is of no use. Instead we seek to assess the internal displacements caused across the breathing cycle. We therefore choose a low-DOF BSpline model.
- Because the implementation of the BRAINSfit module used here expects a 3D volume, we choose a slightly larger grid and make it isotropic, although the 3rd dimension of the grid is not used. We expect the optimizer in the registration to not move outside the slice plane and simply return 0 deformations there.
- An alternative strategy would be to register each image to its immediate previous timepoint, and then integrating the displacements. This has the advantage that nonrigid registration tends to be better for smaller differences, but it bears the risk of accumulating error and hence a lower robustness.
- A related option to improve precision would be to initialize each registration with the result of the previous one. Unfortunately the BRAINSfit registration modules do not (yet) allow to initialize with anything other than an affine.
- The additional information/constraint, that the displacements defined by the series of registrations should describe a smooth motion that is contiguous and differentiable, has not been exploited in this approach. This could be done a posteriori by applying a temporal filter across the transforms or fitting an appropriate model curve to each node displacement.
- Overall strategy:
- select one frame within the breathing cycle as reference frame
- compute non-rigid BSpline registration of all images to the reference frame
- extract the displacements of indiv. grid nodes and plot over time
Procedures
- Phase I: Pilot to determine optimal registration parameters
- load reference image and one moving image from the series
- open Registration : BrainsFit module
- Registration Phases:
- set "reference" fixed and "moving_??" as moving image
- select/check Include BSpline registration phase
- Output Settings:
- select a new transform "Slicer BSpline Transform", rename to "Xf1_moving_??"
- select a new volume "Output Image Volume, rename to "moving_??_Xf1"
- Registration Parameters: increase Number Of Samples to 200,000
- Registration Parameters: set Number Of Grid Subdivisions to 5,5,5
- Leave all other settings at default
- click: Apply; (runtime < 10 sec. on MacPro)
- adjust grid size until registration is acceptable
- you can see the commandline text of the registration performed by opening the Window/Error Log window and clicking on BRAINSfit commandline input
- Phase II: Batch Run
- open a terminal window
- via a TextEditor or prototyping/scripting software (e.g. Matlab), copy and modify the prototype line below, by changing only the moving input image:
/Applications/Slicer36/Slicer3 --launch /Applications/Slicer36/lib/Slicer3/Plugins/BRAINSFit --useBSpline --splineGridSize 5,5,5 --outputVolumePixelType short / --numberOfSamples 200000 --costMetric MMI --fixedVolume Reference/refLung_001.dcm --movingVolume Moving/Moving_001/Moving_001.dcm / --bsplineTransform Xforms/Moving_001_XfBSpl5.tfm --outputVolume MovingResampled/Moving_001_r.nrrd >> Logs/Moving_001_RegLog.txt 2>&1
- replace "/Applications/Slicer36" with your path of 3DSlicer
- create result directories MovingResampled, Logs, Xforms
- note that because input image is DICOM, and images are 2D only, each image of the time series must be in its own directory, otherwise Slicer will read them as a volume.
- paste all commands into a terminal window, or copy into a shell script and execute.
- Phase III: Evaluate Transform files
- upon completion, read the transform files with an editor and extract the displacements of interest
- The ITK transform files describe displacements at the grid nodes, many of which are outside the region of interest. Because BRAINSfit pads the grid with 1 voxel, the grid returned is actually 8x8x8. We use only the plane (*,*,4) for analysis and discard the y-direction displacements, which as expected are all zero.
- for details on the ITK transform format see the FAQ here
- To obtain displacements at arbitrary coordinates, interpolate the transform into a deformation field, e.g. using this module (details FAQ here):
/Applications/Slicer3.6.3/lib/Slicer3/Plugins/BSplineToDeformationField
Registration Results
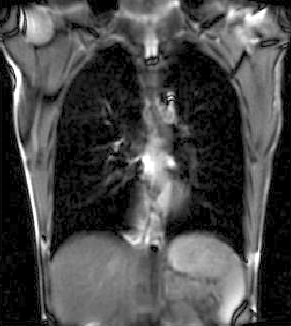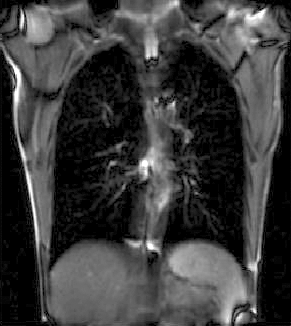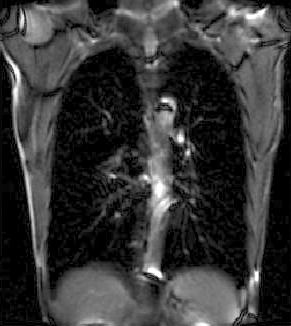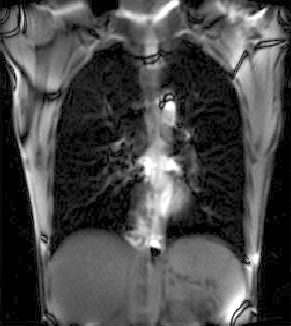
|
|
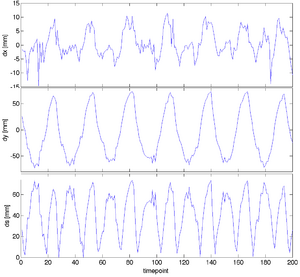
|
| unregistered moving series | moving input after registration (only frames 1-27 shown) | displacements (PDF) |
Download
- Data
- Results
Acknowledgments
Thanks to Amir Owrangi and Dr.Grace Parraga from the Robarts Institute for sharing the registration problem and data.