Difference between revisions of "Slicer3.2:Training"
| Line 33: | Line 33: | ||
| style="background:#D1FFF9; color:black"| [[Media:3DVisualization_SoniaPujol_Munich2008.ppt| Data Loading and Visualization in Slicer3 ]] | | style="background:#D1FFF9; color:black"| [[Media:3DVisualization_SoniaPujol_Munich2008.ppt| Data Loading and Visualization in Slicer3 ]] | ||
| style="background:#D1FFF9; color:black"| [http://www.na-mic.org/Wiki/index.php/Image:SlicerSampleVisualization.tar.gz SlicerSampleVisualization.tar.gz] | | style="background:#D1FFF9; color:black"| [http://www.na-mic.org/Wiki/index.php/Image:SlicerSampleVisualization.tar.gz SlicerSampleVisualization.tar.gz] | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:SceneRestore.png| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:SceneRestore.png|200px|LoadingandVisualization]] |
|- | |- | ||
| style="background:#9BF2C5; color:black" align="Center"| '''1.2''' | | style="background:#9BF2C5; color:black" align="Center"| '''1.2''' | ||
| style="background:#D1FFF9; color:black"| [[Media:AutomaticSegmentation_SoniaPujol_Munich2008.ppt|EM Segmentation Course]]<br>Older materials: [[Media:EMSegTutorial-AHM2008.ppt|1]], [[EMSegmenter| 2]] | | style="background:#D1FFF9; color:black"| [[Media:AutomaticSegmentation_SoniaPujol_Munich2008.ppt|EM Segmentation Course]]<br>Older materials: [[Media:EMSegTutorial-AHM2008.ppt|1]], [[EMSegmenter| 2]] | ||
| style="background:#D1FFF9; color:black"|[[Media:AutomaticSegmentation.tar.gz|AutomaticSegmentation.tar.gz]]<br>Older Materials: [[Media:EMSegTutorial-AHM2008.zip|1]] | | style="background:#D1FFF9; color:black"|[[Media:AutomaticSegmentation.tar.gz|AutomaticSegmentation.tar.gz]]<br>Older Materials: [[Media:EMSegTutorial-AHM2008.zip|1]] | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:EMSegmentation2008.png| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:EMSegmentation2008.png|200px|EM Segmenter]] |
|- | |- | ||
| style="background:#9BF2C5; color:black" align="Center"| '''1.3''' | | style="background:#9BF2C5; color:black" align="Center"| '''1.3''' | ||
| style="background:#D1FFF9; color:black"| Affine and deformable [[Media:Na-MIC-Slicer-Registration.ppt|registration in Slicer 3]] | | style="background:#D1FFF9; color:black"| Affine and deformable [[Media:Na-MIC-Slicer-Registration.ppt|registration in Slicer 3]] | ||
| style="background:#D1FFF9; color:black"| [[Media:SlicerSampleRegistration.tgz|SlicerSampleRegistration.tgz]] | | style="background:#D1FFF9; color:black"| [[Media:SlicerSampleRegistration.tgz|SlicerSampleRegistration.tgz]] | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Na-MIC-Slicer-Registration.png| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Na-MIC-Slicer-Registration.png|200px|SlicerRegistration]] |
|- | |- | ||
| style="background:#9BF2C5; color:black" align="Center"| '''1.4''' | | style="background:#9BF2C5; color:black" align="Center"| '''1.4''' | ||
| Line 50: | Line 50: | ||
See [http://slicer.org/slicerWiki/index.php/Slicer3:DTMRI '''here'''] for background info. | See [http://slicer.org/slicerWiki/index.php/Slicer3:DTMRI '''here'''] for background info. | ||
| style="background:#D1FFF9; color:black"|[[Media:Slicer3-diffusion-Tutorial material.zip|Slicer3-diffusion-Tutorial material.zip]] | | style="background:#D1FFF9; color:black"|[[Media:Slicer3-diffusion-Tutorial material.zip|Slicer3-diffusion-Tutorial material.zip]] | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Line-glyph-tracts.jpg| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Line-glyph-tracts.jpg|200px|Glyphs and tracts]] |
|- | |- | ||
| style="background:#9BF2C5; color:black" align="Center"| '''1.5''' | | style="background:#9BF2C5; color:black" align="Center"| '''1.5''' | ||
| Line 56: | Line 56: | ||
This tutorial consists of two MR scans of a patient with meningioma. | This tutorial consists of two MR scans of a patient with meningioma. | ||
| style="background:#D1FFF9; color:black"|[[Media:TumorGrowth-Tutorial-Data.xcat|TumorGrowth-Tutorial-Data.xcat ]] | | style="background:#D1FFF9; color:black"|[[Media:TumorGrowth-Tutorial-Data.xcat|TumorGrowth-Tutorial-Data.xcat ]] | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:TumorGrowth-Tutorial-Image.png| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:TumorGrowth-Tutorial-Image.png|200px|Meningioma Case]] |
|- | |- | ||
| style="background:#8EDEB5; color:black" align="Center"| '''2.1''' | | style="background:#8EDEB5; color:black" align="Center"| '''2.1''' | ||
| Line 62: | Line 62: | ||
This tutorial is intended for engineers and scientists who want to integrate external programs with Slicer3. | This tutorial is intended for engineers and scientists who want to integrate external programs with Slicer3. | ||
| style="background:#D1FFF9; color:black"| [[Media:HelloWorld.zip|HelloWorld.zip]] | | style="background:#D1FFF9; color:black"| [[Media:HelloWorld.zip|HelloWorld.zip]] | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:HelloWorld.png| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:HelloWorld.png|200px|Plug-In]] |
|- | |- | ||
| style="background:#8EDEB5; color:black" align="Center"| '''2.2''' | | style="background:#8EDEB5; color:black" align="Center"| '''2.2''' | ||
| Line 69: | Line 69: | ||
This tutorial takes the trainee through a complete workup for neurosurgical patient specific mapping | This tutorial takes the trainee through a complete workup for neurosurgical patient specific mapping | ||
| style="background:#D1FFF9; color:black"|[[Media:NeurosurgicalPlanningTutorialData.zip|NeurosurgicalPlanningTutorialData.zip]] | | style="background:#D1FFF9; color:black"|[[Media:NeurosurgicalPlanningTutorialData.zip|NeurosurgicalPlanningTutorialData.zip]] | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:NeurosurgicalPlanningOverview.png| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:NeurosurgicalPlanningOverview.png|200px|Neurosurgical Planning Overview]] |
|- | |- | ||
| style="background:#72b291; color:black" align="Center"| '''3.1''' | | style="background:#72b291; color:black" align="Center"| '''3.1''' | ||
| Line 75: | Line 75: | ||
This tutorial is intended for engineers and scientists who want to use Slicer 3 for IGT research. | This tutorial is intended for engineers and scientists who want to use Slicer 3 for IGT research. | ||
| style="background:#D1FFF9; color:black"| | | style="background:#D1FFF9; color:black"| | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Slicer IGTL NITRobot.jpg| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Slicer IGTL NITRobot.jpg|200px|Slicer with robots]] |
|- | |- | ||
| style="background:#72b291; color:black" align="Center"| '''3.2''' | | style="background:#72b291; color:black" align="Center"| '''3.2''' | ||
| Line 81: | Line 81: | ||
This three-dimensional brain atlas dataset, derived from a volumetric, whole-head MRI scan, contains the original MRI-scan, a complete set of label maps, three-dimensional reconstructions (200+ structures) and a tutorial. See [http://www.spl.harvard.edu/pages/Special:PubDB_View?dspaceid=1265 '''here'''] for more information. | This three-dimensional brain atlas dataset, derived from a volumetric, whole-head MRI scan, contains the original MRI-scan, a complete set of label maps, three-dimensional reconstructions (200+ structures) and a tutorial. See [http://www.spl.harvard.edu/pages/Special:PubDB_View?dspaceid=1265 '''here'''] for more information. | ||
| style="background:#D1FFF9; color:black"|[http://www.spl.harvard.edu/pages/Special:PubDB_View?dspaceid=1265 SPL-PNL Brain Atlas] | | style="background:#D1FFF9; color:black"|[http://www.spl.harvard.edu/pages/Special:PubDB_View?dspaceid=1265 SPL-PNL Brain Atlas] | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Atlas.png| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Atlas.png|200px|Brain Atlas]] |
|- | |- | ||
| style="background:#72b291; color:black" align="Center"| '''3.3''' | | style="background:#72b291; color:black" align="Center"| '''3.3''' | ||
| Line 87: | Line 87: | ||
This three-dimensional abdominal atlas was derived from a computed tomography (CT) scan, using semi-automated image segmentation and three-dimensional reconstruction techniques. The dataset contains the original CT scan, detailed label maps, three-dimensional reconstructions and a tutorial. See [http://www.spl.harvard.edu/pages/Special:PubDB_View?dspaceid=1266 '''here'''] for more information. | This three-dimensional abdominal atlas was derived from a computed tomography (CT) scan, using semi-automated image segmentation and three-dimensional reconstruction techniques. The dataset contains the original CT scan, detailed label maps, three-dimensional reconstructions and a tutorial. See [http://www.spl.harvard.edu/pages/Special:PubDB_View?dspaceid=1266 '''here'''] for more information. | ||
| style="background:#D1FFF9; color:black"|[http://www.spl.harvard.edu/pages/Special:PubDB_View?dspaceid=1266 SPL Abdominal Atlas] | | style="background:#D1FFF9; color:black"|[http://www.spl.harvard.edu/pages/Special:PubDB_View?dspaceid=1266 SPL Abdominal Atlas] | ||
| − | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Abdominal-Atlas-3Dsnapshot.png| | + | | style="background:#C3D1C3; color:black" align="Center"| [[Image:Abdominal-Atlas-3Dsnapshot.png|200px|Abdominal Atlas]] |
|} | |} | ||
Revision as of 14:07, 8 August 2008
Home < Slicer3.2:TrainingWelcome to the 3D Slicer3.2 User Training 101
This page is under construction The information on this page applies to 3D Slicer version 3. If you are looking for materials about Slicer version 2, please go to the following location.
|
Slicer Logo | |
Software Installation
The Slicer download page contains links for downloading the different versions of Slicer 3.
Training Compendium
| Category/# | Tutorial | Sample Data | Image |
| 1.1 | Data Loading and Visualization in Slicer3 | SlicerSampleVisualization.tar.gz | 
|
| 1.2 | EM Segmentation Course Older materials: 1, 2 |
AutomaticSegmentation.tar.gz Older Materials: 1 |

|
| 1.3 | Affine and deformable registration in Slicer 3 | SlicerSampleRegistration.tgz | 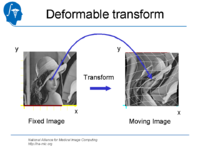
|
| 1.4 | Processing of DWI and DTI data in Slicer3- WARNING UNDER CONSTRUCTION NOT READY FOR USE earlier version This tutorial takes the trainee through the slicer3 diffusion display and processing options. See here for background info. |
Slicer3-diffusion-Tutorial material.zip | 
|
| 1.5 | Detecting subtle change in pathology This tutorial consists of two MR scans of a patient with meningioma. |
TumorGrowth-Tutorial-Data.xcat | 
|
| 2.1 | Plug-ins for Slicer3: Course for Developers
This tutorial is intended for engineers and scientists who want to integrate external programs with Slicer3. |
HelloWorld.zip | 
|
| 2.2 | Image Guided Therapy Planning Tutorial See here for background info. This tutorial takes the trainee through a complete workup for neurosurgical patient specific mapping |
NeurosurgicalPlanningTutorialData.zip | 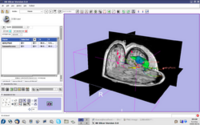
|
| 3.1 | Slicer3 as a research tool for image guided therapy research (IGT)
This tutorial is intended for engineers and scientists who want to use Slicer 3 for IGT research. |

| |
| 3.2 | SPL-PNL Brain Atlas
This three-dimensional brain atlas dataset, derived from a volumetric, whole-head MRI scan, contains the original MRI-scan, a complete set of label maps, three-dimensional reconstructions (200+ structures) and a tutorial. See here for more information. |
SPL-PNL Brain Atlas | 
|
| 3.3 | SPL Abdominal Atlas
This three-dimensional abdominal atlas was derived from a computed tomography (CT) scan, using semi-automated image segmentation and three-dimensional reconstruction techniques. The dataset contains the original CT scan, detailed label maps, three-dimensional reconstructions and a tutorial. See here for more information. |
SPL Abdominal Atlas | 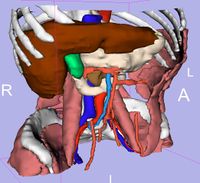
|
- Category 1 = Basic functionality
- Category 2 = Advanced functionality
- Category 3 = Specialized application packages
Additional Materials
For a variety of data sets for downloading, check the following link.
Back to Training:Main
