NA-MIC NCBC Collaboration:3D+t Cells Lineage:GoFigure
 Return to Project Week Main Page |
Key Investigators
- Sean Megason (Caltech)
- Alex. Gouaillard (Caltech)
- Titus Brown (Caltech)
- Yumin Yuan (Kitware)
- Luis Ibanez (Kitware)
Objective
- Distributed Image database
- Very large datasets (200+ Gb)
- Segmentation of cells in 2D+t, 3D and 3D+t datasets
- New Data structure for 2-Manifolds (itkQE)
- Cross-platform GUI
- investigate Collaboration with NAMIC
- hire graduate students for the project.
Approach, Plan
- recode / wrap segmentation algos in VTK / ITK
- test different segmentations algorithms.
- plug in other VTK / ITK segmentation filters.
- try to parallelise dataset access through DB.
- investgate KWWidgets for GUI.
- finish porting QE in ITK.
Progress
- IN PROGRESS
- DONE
- IN PROGRESS
- .
- DONE
- DONE
others
- No candidate (sic)
- Code Coverage / Memory Checking + dashboard
Scientific Background
Imaging for System Biology
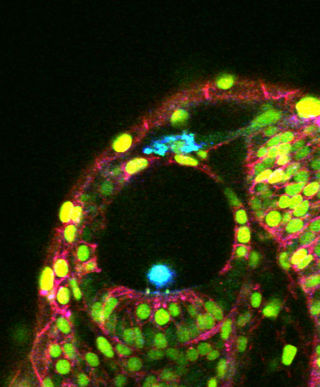
Imaging is becoming a very important approach for doing systems biology because of its unique ability to capture quantitative, longitudinal data at cellular resolution from living organisms. However, the use of imaging for systems biology presents unique and significant challenges for image analysis. We have recently established a Center of Excellence in Genomic Science at Caltech to develop the technology (imaging hardware, image analysis software, and genetic/molecular tools) to develop an imaging based platform for deciphering zebrafish embryogenesis.
In Toto imaging
In Toto Genomic Analysis of Vertebrate Development
This Center of Excellence in Genomic Science (CEGS) assembles a multidisciplinary group of Caltech investigators, Sean Megason, Niles Pierce, Scott Fraser and Marianne Bronner-Fraser, to develop innovative technologies with the goal of imaging and mutating every developmentally important vertebrate gene. Novel "in toto imaging" tools will make it possible to analyze gene expression and function in developing vertebrate embryos in time and space, digitizing in vivo data in a systematic, high-throughput, and quantitative fashion.
Key technologies will be developed and tested in the zebrafish embryo since it is ideal for both imaging and genetics. We are generating FlipTraps to fluorescently mark protein expression and gene function on a genome wide scale, and developing new techniques for in situ hybridization using HCR that permit simultaneous analysis of multiple marker genes. In toto imaging will be used to digitize this marked data quantitatively at cellular resolution throughout embryogenesis. The vast amount of quantitative, in vivo data obtained through this project will serve as the basis for creating a "digital fish" that models the molecular and cellular orchestra that transforms an egg into an embryo.
GoFigure
GoFigure is an image analysis software package specifically designed for visualizing, tracking, and analyzing cells in multidimensional confocal/two-photon image sets. Eventually GoFigure will be able to analyze time-lapse series of confocal image stacks (xyzt) and to automatically segment and track cells in 2D, 2D+time, 3D, 3D+time, and 3D+time+cell division.
GoFigure now runs on the Microsoft Windows platform. GoFigure utilizes a MySql database back-end so it can handle extremely large image sets.
Open Source and Open Software
The Beta version of GoFigure is still only distributed on-demand, the project embrace the OpenSource and OpenData philosophy. We want the software and the data not only to be at the disposition of the researchers but we also want the researchers to be able to make their own modifications and enhancements to it. the transition phase from the Alpha version (prototype) to a full open source software is under work. The actual Alpha version is base on MFC and VTK for the vizualization. The Beta version will be based on the following NA-MIC kits: VTK, ITK, CMake, CTest, CPack and on the following OpenSource software and technologies: svn, track, wiki, mailing list, ... We have postdoctoral positions in the Department of Systems Biology at Harvard and the Departments of Biology and Applied Math at Caltech to fill related to GoFigure development.
Prototype Code in the NAMIC Sandbox
Prototype code for the segmentation of cells from confocal microscopy datasets will be stored at:
http://www.na-mic.org/svn/NAMICSandBox/trunk/CellLineageSegmentation/
Progress before project week:
- beta version available
- itkQE's core in review for integration in ITK
- cmake-compliant configuration/compilation
- packaging (cpack)
- Dashboard (dart2)