Difference between revisions of "2010 Summer Project Week MRSI module and SIVIC interface"
(Created page with '__NOTOC__ <gallery> Image:PW-MIT2010.png|Projects List Image:Spectral fitting mrs.png|Fitting metabolite models to the MRS signal of tumorou…') |
(No difference)
|
Revision as of 18:42, 31 May 2010
Home < 2010 Summer Project Week MRSI module and SIVIC interfaceKey Investigators
- MIT: Bjoern Menze, M Phothilimthana, Polina Golland
- UCSF: Olson Beck, Jason Crane (Nelson Lab)
Project
Objective
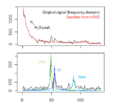
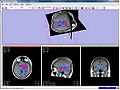
Magnetic resonance spectroscopic imaging (MRSI) is a non-invasive diagnostic method used to determine the relative abundance of specific metabolites at arbitrary locations in vivo. Certain diseases -- such as tumors in brain, breast and prostate -- can be can be associated with characteristic changes in the metabolic level.
The objective of the current project is to develop a module proving the means for the processing and visualization of MRSI -- and thus for a joint analysis of magnetic resonance spectroscopic images together with other imaging modalities -- in Slicer.
Approach, Plan
Spectral fitting routines have been implemented, using a HSVD filter for water peak removal and baseline estimation, and a constrained non-linear least squares optimization for the fit of the resonance line models. Current fitting routines require, however, several external software libraries not to be distributed, installed and used easily.
We want to integrate the currently available MRSI quantitation routines with the SIVIC Slicer interface developed at the UCSF.