Difference between revisions of "2010 Winter Project Week The Vascular Modeling Toolkit in 3D Slicer"
(Created page with '__NOTOC__ <gallery> Image:PW-SLC2010.png|Projects List Image:genuFAp.jpg|Scatter plot of the original FA data through the genu of the corpus…') |
|||
| (7 intermediate revisions by 2 users not shown) | |||
| Line 2: | Line 2: | ||
<gallery> | <gallery> | ||
Image:PW-SLC2010.png|[[2010_Winter_Project_Week#Projects|Projects List]] | Image:PW-SLC2010.png|[[2010_Winter_Project_Week#Projects|Projects List]] | ||
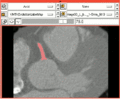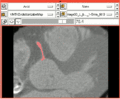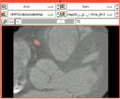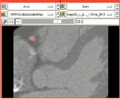
| − | Image: | + | Image:Vmtkafterevolution.png|A coronary artery segmented using VMTK in 3D Slicer. |
| − | Image: | + | Image:Vmtkcloseupvoronoicenterlinewithreference.png|Close-up showing the corresponding Voronoi diagram and centerline. |
| + | Image:Vmtkafterevolutionanim.gif|An over-layed label map showing the segmentation in 2D. | ||
| + | Image:Pokal.gif|End-User Tutorial Contest Winner | ||
</gallery> | </gallery> | ||
==Key Investigators== | ==Key Investigators== | ||
| − | * [[User:haehn |Daniel Haehn]] | + | * [[User:haehn |Daniel Haehn]], Student of Medical Informatics, University of Heidelberg, Germany |
| + | * Steve Pieper, Isomics, Inc. | ||
| + | * Luca Antiga, Mario Negri Institute, Italy | ||
<div style="margin: 20px;"> | <div style="margin: 20px;"> | ||
| Line 17: | Line 21: | ||
| + | The official project page: http://www.vmtk.org/Main/VmtkIn3DSlicer | ||
| Line 24: | Line 29: | ||
<h3>Approach, Plan</h3> | <h3>Approach, Plan</h3> | ||
| + | To provide VMTK functionality in 3D Slicer the ''vtkVmtk'' library has to be available as an add-on. With the connection of VMTK and 3D Slicer processing pipelines | ||
| + | between VMTK code and other algorithms can be established. | ||
| + | |||
| + | '''Plan for project week:''' Clean-up of existing VMTK in 3D Slicer code, maybe adding additional functionality, investigating GWE possibilities.. | ||
</div> | </div> | ||
| Line 31: | Line 40: | ||
<h3>Progress</h3> | <h3>Progress</h3> | ||
| + | The VMTK library and modules providing selected functionality are available as ''3D Slicer extensions''. This enables a flexible and convenient way for distribution and installation. | ||
| + | |||
| + | |||
| + | The following extensions are available: | ||
| + | * '''VmtkSlicerModule''' - a gui-less module providing the latest VMTK library (base installation) | ||
| + | * '''VMTKLevelSetSegmentation''' - providing the VMTK level-set segmentation process including different algorithms for initialization and evolution | ||
| + | * '''VMTKEasyLevelSetSegmentation''' - an easier interface to level-set segmentation with selected algorithms | ||
| + | * '''VMTKVesselEnhancement''' - Vesselness filtering to enhance tubular structures | ||
| + | * '''VMTKCenterlines''' - Centerline computation of polydata models | ||
| + | |||
| + | |||
| + | The modules have been successfully applied to segmentation problems (e.g. Coronary Artery Centerline Extraction - see [http://wiki.slicer.org/slicerWiki/index.php/Modules:VMTK_in_3D_Slicer_Tutorial:_Coronary_Artery_Centerline_Extraction Tutorial]). | ||
</div> | </div> | ||
</div> | </div> | ||
| + | <div style="width: 97%; float: left;"> | ||
| + | ==Project Week Results== | ||
| + | * VMTK in 3D Slicer ready to use with Grid Wizard Enterprise (http://www.gridwizardenterprise.org) | ||
| + | * S2EL files to access the VMTK functionality. (Import of the logic classes) | ||
| + | * '''For example Frangi's Vesselness:''' | ||
| + | <pre> | ||
| + | |||
| + | ${SIGMA_MIN}=$range(0.1,1.0,0.1) | ||
| + | ${SIGMA_MAX}=$range(1.0,4.0,0.5) | ||
| + | ${SIGMA_STEPS}=$const(10) | ||
| + | ${ALPHA}=$range(0.1,1.0,0.1) | ||
| + | ${BETA}=$range(5.0,10.0,1.0) | ||
| + | ${GAMMA}=$range(5.0,10.0,1.0) | ||
| + | ${OUTPUT}=$const(/home/hype/gwe/results/frangi_out-${SIGMA_MIN}-${SIGMA_MAX}-N${SIGMA_STEPS}-${ALPHA}-${BETA}-${GAMMA}.nrrd) | ||
| + | |||
| + | ${SLICER_HOME}/Slicer3 --no_splash --evalpython | ||
| + | "import sys; | ||
| + | from Slicer import slicer; | ||
| + | sys.path.append(str(slicer.Application.GetExtensionsInstallPath())+'/'+str(slicer.Application.GetSvnRevision())+'/VMTKVesselEnhancement/VMTKVesselEnhancement'); | ||
| + | from SlicerVMTKVesselEnhancementGUI import *; | ||
| + | hiddengui = VMTKVesselEnhancement(); | ||
| + | from SlicerVMTKVesselEnhancementLogic import *; | ||
| + | vesselness=SlicerVMTKVesselEnhancementLogic(hiddengui); | ||
| + | |||
| + | volNode=slicer.VolumesGUI.GetLogic().AddArchetypeVolume('/home/hype/gwe/liver.nrrd','Liver',0); | ||
| + | matrix = slicer.vtkMatrix4x4(); | ||
| + | volNode.GetIJKToRASMatrix(matrix); | ||
| + | |||
| + | outVolumeData = vesselness.ApplyFrangiVesselness(volNode.GetImageData(),${SIGMA_MIN},${SIGMA_MAX},${SIGMA_STEPS},${ALPHA},${BETA},${GAMMA}); | ||
| + | outputNode=slicer.MRMLScene.CreateNodeByClass('vtkMRMLScalarVolumeNode'); | ||
| + | volumeNode=slicer.MRMLScene.AddNode(outputNode); | ||
| + | volumeNode.SetAndObserveImageData(outVolumeData); | ||
| + | volumeNode.SetIJKToRASMatrix(matrix); | ||
| + | volumeNode.SetModifiedSinceRead(1); | ||
| + | slicer.VolumesGUI.GetLogic().SaveArchetypeVolume(${OUTPUT},volumeNode);" | ||
| + | |||
| + | </pre> | ||
| + | </div> | ||
<div style="width: 97%; float: left;"> | <div style="width: 97%; float: left;"> | ||
==References== | ==References== | ||
| + | *Antiga L, Piccinelli M, Botti L, EneIordache B, Remuzzi A, Steinmann DA. (2008) An imagebased modeling framework for patientspecific computational hemodynamics. Med Biol Eng Comput 46(11):10971112 | ||
| + | *Antiga L, Steinman DA (2008) The Vascular Modeling Toolkit. http://www.vmtk.org/ | ||
| + | *Hähn D (2009) [http://www.spl.harvard.edu/publications/bitstream/download/4629 Integration of The Vascular Modeling Toolkit in 3D Slicer]. Student Research Project, SPL. | ||
| + | *Piccinelli M, Veneziani A, Steinman DA, Remuzzi A, Antiga L (2009) A framework for geometric analysis of vascular structures: applications to cerebral aneurysms. IEEE Trans Med Imaging. In press. | ||
</div> | </div> | ||
Latest revision as of 17:12, 8 January 2010
Home < 2010 Winter Project Week The Vascular Modeling Toolkit in 3D Slicer
Key Investigators
- Daniel Haehn, Student of Medical Informatics, University of Heidelberg, Germany
- Steve Pieper, Isomics, Inc.
- Luca Antiga, Mario Negri Institute, Italy
Objective
The Vascular Modeling Toolkit (VMTK) is a collection of libraries and tools for 3D reconstruction, geometric analysis, mesh generation and surface data analysis for image-based modeling of blood vessels. It should be very interesting to offer such techniques in 3D Slicer.
The official project page: http://www.vmtk.org/Main/VmtkIn3DSlicer
Approach, Plan
To provide VMTK functionality in 3D Slicer the vtkVmtk library has to be available as an add-on. With the connection of VMTK and 3D Slicer processing pipelines between VMTK code and other algorithms can be established.
Plan for project week: Clean-up of existing VMTK in 3D Slicer code, maybe adding additional functionality, investigating GWE possibilities..
Progress
The VMTK library and modules providing selected functionality are available as 3D Slicer extensions. This enables a flexible and convenient way for distribution and installation.
The following extensions are available:
- VmtkSlicerModule - a gui-less module providing the latest VMTK library (base installation)
- VMTKLevelSetSegmentation - providing the VMTK level-set segmentation process including different algorithms for initialization and evolution
- VMTKEasyLevelSetSegmentation - an easier interface to level-set segmentation with selected algorithms
- VMTKVesselEnhancement - Vesselness filtering to enhance tubular structures
- VMTKCenterlines - Centerline computation of polydata models
The modules have been successfully applied to segmentation problems (e.g. Coronary Artery Centerline Extraction - see Tutorial).
Project Week Results
- VMTK in 3D Slicer ready to use with Grid Wizard Enterprise (http://www.gridwizardenterprise.org)
- S2EL files to access the VMTK functionality. (Import of the logic classes)
- For example Frangi's Vesselness:
${SIGMA_MIN}=$range(0.1,1.0,0.1)
${SIGMA_MAX}=$range(1.0,4.0,0.5)
${SIGMA_STEPS}=$const(10)
${ALPHA}=$range(0.1,1.0,0.1)
${BETA}=$range(5.0,10.0,1.0)
${GAMMA}=$range(5.0,10.0,1.0)
${OUTPUT}=$const(/home/hype/gwe/results/frangi_out-${SIGMA_MIN}-${SIGMA_MAX}-N${SIGMA_STEPS}-${ALPHA}-${BETA}-${GAMMA}.nrrd)
${SLICER_HOME}/Slicer3 --no_splash --evalpython
"import sys;
from Slicer import slicer;
sys.path.append(str(slicer.Application.GetExtensionsInstallPath())+'/'+str(slicer.Application.GetSvnRevision())+'/VMTKVesselEnhancement/VMTKVesselEnhancement');
from SlicerVMTKVesselEnhancementGUI import *;
hiddengui = VMTKVesselEnhancement();
from SlicerVMTKVesselEnhancementLogic import *;
vesselness=SlicerVMTKVesselEnhancementLogic(hiddengui);
volNode=slicer.VolumesGUI.GetLogic().AddArchetypeVolume('/home/hype/gwe/liver.nrrd','Liver',0);
matrix = slicer.vtkMatrix4x4();
volNode.GetIJKToRASMatrix(matrix);
outVolumeData = vesselness.ApplyFrangiVesselness(volNode.GetImageData(),${SIGMA_MIN},${SIGMA_MAX},${SIGMA_STEPS},${ALPHA},${BETA},${GAMMA});
outputNode=slicer.MRMLScene.CreateNodeByClass('vtkMRMLScalarVolumeNode');
volumeNode=slicer.MRMLScene.AddNode(outputNode);
volumeNode.SetAndObserveImageData(outVolumeData);
volumeNode.SetIJKToRASMatrix(matrix);
volumeNode.SetModifiedSinceRead(1);
slicer.VolumesGUI.GetLogic().SaveArchetypeVolume(${OUTPUT},volumeNode);"
References
- Antiga L, Piccinelli M, Botti L, EneIordache B, Remuzzi A, Steinmann DA. (2008) An imagebased modeling framework for patientspecific computational hemodynamics. Med Biol Eng Comput 46(11):10971112
- Antiga L, Steinman DA (2008) The Vascular Modeling Toolkit. http://www.vmtk.org/
- Hähn D (2009) Integration of The Vascular Modeling Toolkit in 3D Slicer. Student Research Project, SPL.
- Piccinelli M, Veneziani A, Steinman DA, Remuzzi A, Antiga L (2009) A framework for geometric analysis of vascular structures: applications to cerebral aneurysms. IEEE Trans Med Imaging. In press.