2014 Winter Project Week:SubjectHierarchy
From NAMIC Wiki
Home < 2014 Winter Project Week:SubjectHierarchy
Key Investigators
- Queen's: Csaba Pinter, Andras Lasso
- Isomics: Steve Pieper
- Kitware: Jean-Cristophe Fillion-Robin
Project Description
Objective
- Review the current state of Subject Hierarchy
- Discuss features to develop and things to change
- Create roadmap for the integration
Approach, Plan
Progress
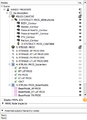
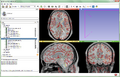
- Subject hierarchy enhanced to better support handling Huntington's longitudinal data, video
- Volumes subject hierarchy plugin handles labelmap and proper show/hide of volumes in the slice viewers
- Subject hierarchy module and mechanism presented in the breakot session, a lot of valuable feedback was given (see below)
Received feedback
- DICOM back into SH? Derived vtkMRMLDICOMSubjectHierarchyNode class? For tags, UID, maybe levels?
- DICOM connections
- "All DICOM all the time" (@QIICR) means for import/export on disc, not in memory
- Tree vs DAG: Support DAGs? How to display them? "Infinite trees"?
- Update referenced UIDs and references when reparenting and the connections are known (plugins reparent method)
- Look at BioImageSuite (by Xenios) - User manual chapter 15 (page 193)
- Connector lines in the tree are colored (indicate status of registration?)
- Export
- Override the selection made by the user to the most sensible one (if they select a contour, the RTSS is also selected)
- Determine mandatory tags for export
- Stored in the plugins?
- Maybe in XML or JSON templates that are read by the plugins?
- Collect tags from plugin dependencies
- Stored in the plugins?
- Use DICOM header widget with some changes (ctkDICOMObjectModel + ListWidget)
- Similar modalities to export (legacy, multiframe, enhanced) - separate plugins?
- Explicit representation of DICOM tags in Slicer in separate class?
- Ask for mandatory DICOM tags and connections (e.g. after conversion, resample, manual plugin selection)
- Use cases
- Neuro
- Multiple modalities (different MR types)
- Display tractography
- Register volumes and indicate the registration status in the tree (in the icons)
- Data probe custom readout (e.g. dose)
- DICOM slice offset into the slice views
- Displayable manager customizable behavior
- Scalar volume types e.g. multi-component vector volumes
- Volume rendering transfer functions
- Neuro
- UIDs
- "2.25.hash" DICOM UIDs will be generated as new UIDs (perhaps even instead of MRML IDs in the future)
- What happens to the UIDs if we modify a node?
- Create new SH node and move the data there?
- MIDAS url should be a SH storage node property? (Julien)
- Plugins
- Give confidence value of -1 if the node cannot be handled for sure?
- Per-structure labelmaps
- More abstraction: Qt, JavaScript etc. -> presentation managers
- Icon could be in the MRML node as a resource (string)
- Move core plugins into their corresponding modules after integration (Volumes, Transforms, Markups, etc.)
- Create dummy parents for loaded node if there is no parent (Study, Patient too)
- 'Unclaimed' section on the bottom of the tree instead of potential list? (Jim)
- Drag&drop would be difficult if there is a lot of nodes in the tree already
- Generate thumbnails as icons for volumes
- vtkSubjectHierarchyConstants -> vtkMRMLSubjectHierarchyConstants?
- qMRMLSortFilterPotentialSubjectHierarchyProxyModel be merged into qMRMLSortFilterSubjectHierarhcyProxyModel?
- Maybe a Q_PROPERTY controls their different behavior?
- Propagating / Overwriting attributes
- In the model, for example for "height" of an item
- Select and drag&drop multiple nodes from the potential list