Algorithm:UNC:Old
UNC Algorithms page
Contents
Diffusion Tensor Imaging
DT-MRI provides a new method to investigate the geometry and properties of white matter in-vivo. Major goals of our research in DT-MRI analysis include tract-based regions of interest for statistics, population based analysis, and understanding the influence of noise on statistical measures.
Quantitative Analysis of Fiber Tract Bundles
DT-MRI tractography can be used as a coordinate system for computing statistics of diffusion tensor data. The quantitative analysis of diffusion tensors takes into account the space of tensor measurements using a nonlinear Riemannian symmetric space framework. Tracts of interest are represented as a medial spline attributed with cross-sectional statistics.
Description - Publications - Software
Population Analysis from Deformable Registration
Analysis of populations of diffusion images typically requires time-consuming manual segmentation of structures of interest to obtain correspondance for statistics. This project uses non-rigid registration of DTI images to produce a common coordinate system for hypothesis testing of diffusion properties.
New:
- Application to PNL data
- Command line DTI tools available as part of UNC NeuroLib
Description - Publications - Software
Influence of Imaging Noise on DTI Statistics (Collaboration with Utah)
Clinical acquisition of diffusion weighted images with high signal to noise ratio remains a challenge. The goal of this project is to understand the impact of MR noise on quantiative statistics of diffusion properties such as anisotropy measures, trace, etc.
New:
- Casey Goodlett, P. Thomas Fletcher, Weili Lin, Guido Gerig. Quantification of measurement error in DTI: Theoretical predictions and validation. To Appear MICCAI 2007
Description - Publications - Software
Shape Analysis of Brain Structures Across Groups
Shape analysis has become of increasing relevance to the neuroimaging community due to its potential to precisely locate morphological changes between healthy and pathological structures. This project focuses on developing novel methodology and a comprehensive set of tools for the computation of 3D structural statistical shape analysis. There are several open problems in this area, ranging from multi-object analysis, enhanced shape correspondence to statistical analysis of shape with clinical covariates.
UNC Shape Analysis Framework using SPHARM-PDM
|
The UNC shape analysis is based on an analysis framework of objects with spherical topology, described mainly by sampled spherical harmonics SPHARM-PDM. The input of the shape analysis framework is a set of binary segmentations of a single brain structure, such as the hippocampus or caudate. These segmentations are converted into a shape description (SPHARM) with correspondence and analyzed via Hotelling T^2 two sample metric. More... New:
|
Population Based Correspondence
We are developing methodology to automatically find dense point correspondences between a collection of polygonal genus 0 meshes. The advantage of this method is independence from indivisual templates, as well as enhanced modeling properties. The method is based on minimizing a cost function that describes the goodness of correspondence. Apart from a cost function derived from the description length of the model, we also employ a cost function working with arbitrary local features. We extended the original methods to use surface curvature measurements, which are independent to differences of object aligment. More...
New:
- Software available as part of UNC Neurolib open source (website)
- Evaluation on lateral ventricles, hippocampi, caudates, striatum, femural bone. Outperforms standard MDL on complex structures.
- Submission to MICCAI 2007 conference
- Tobias Heimann, I. Oguz, I. Wolf, M. Styner, HP. Meinzer. Implementing the Automatic Generation of 3D Statistical Shape Models with ITK. Accepted to MICCAI 2006 Open Source Workshop. More...
Description - Publications - Software
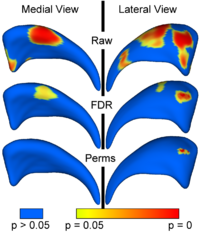
Local Statistical Analysis via Permutation Tests
We have further developed a set of statistical testing methods that allow the analysis of local shape differences using the Hotelling T 2 two sample metric. Permutatioin tests are employed for the computation of statistical p-values, both raw and corrected for multiple comparisons. Resulting significance maps are easily visualized. Additional visualization of the group tests are provided via mean difference magnitude and vector maps, as well as maps of the group covariance information. Ongoing research focuses on incorporating covariates such as clinical scores into the testing scheme. More...
New:
- Available as part of Shape Analysis Toolset in UNC Neurolib open source (download).
- M. Styner, I. Oguz, S. Xu, C. Brechbuehler, D. Pantazis, J. Levitt, M. Shenton, G. Gerig. Framework for the Statistical Shape Analysis of Brain Structures using SPHARM-PDM. Accepted to MICCAI 2006 Open Source Workshop. More...
Description - Publications - Software
Collaborations with other groups in NAMIC
- Algorithms:
- Shape Analysis
- Joint pipeline I/O formulation and development with Kitware (Brad Davis, Jim Miller) and MIT (Polina Golland)
- Use of UNC statistical analysis for spherical wavelet shape with GeorgiaTech (Delphine Nain) and Utah (Tom Fletcher)
- Use of UNC statistical analysis for combined multi-object correspondence establishment with Utah (Josh Cates, Tom Fletcher)
- DTI
- Statistics of tensors and noise in diffusion weighted imaging with Utah (Tom Fletcher)
- Shape Analysis
- Clinical:
- Collaboration with Harvard on shape analysis and DTI analysis.