DBP3:MGH
Back to NA-MIC DBPs | NA-MIC Cores
Introduction
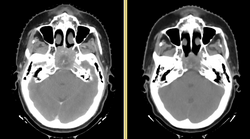
Head and neck cancers account for about 60,000 new cancer cases per year and represent about 6% of all cancers in the United States [1]. These cancers are treated by a combination of chemotherapy, radiotherapy, and surgery. The five-year survival is approximately 50%. During a six-week regimen of radiotherapy, head and neck cancer patients often exhibit anatomic changes that affect their treatment. These changes include tumor regression or growth, changes in lymph node size, and changes in air cavities. Uncorrected, these changes can increase the risk of treatment complications or reduce treatment efficacy.
Adaptive radiotherapy addresses the problem of anatomic change by incrementally adjusting the radiotherapy plan, and is a prime example of personalized medicine. A mid-treatment adjustment is complex: it requires a new CT image, image segmentation, deformable registration, and mapping of the previously delivered dose onto the new image. This project proposes to use the NA-MIC Kit to develop a simple, practical workflow for achieving adaptive radiotherapy which can be applied on a case-by-case basis.
For details on the background and goals, please see the project web page.
For a more extensive wishlist, see the planning meeting notes
Project Goals
State of the Art: 2010
Research Plan
The overall goal of the project is to improve the capabilities for
- Goal: Scientific evidence for or against adaptive proton-beam radiotherapy in the base of skull region
- Require: Update literature review
- Recommend: Study to quantify geometry of anatomic differences
- Require: Comparison study of delivered vs. planned dose
- Require: Comparison study of delivered vs. adaptive dose
- Wishlist: Comparison study of IMRT vs. protons
Engineering
See: http://www.na-mic.org/Wiki/index.php/EngineeringRetreat2010
- Pushing CMake module bug fixes upstream
- Window/level slider bug 1047
- Add mode compositing glitches 1048 1049
Image Registration
- Goal: Multiple slicer plugins for registration
- Recommend: Experimental evaluation of existing registration methods
- Require: Interactive registration (landmark splines)
- Recommend: Regularized B-splines
- Recommend: B-splines with landmark constraints
- Recommend: B-splines with surface constraints
- Wishlist: Parameter-free registration
- Wishlist: Image-free surface registration
- Wishlist: Parallel optimization
Image Segmentation
- Goal: Slicer plugin for automatic segmentation
- Require: Contour propagation for intra-subject segmentation
- Recommend: Automatic segmentation (atlas-based) for inter-subject segmentation
- Recommend: Automatic segmentation (model-based) for inter-subject segmentation
- Wishlist: Contour interpolation methods for interactive segmentation
Radiotherapy Workflow
- Goal: Reasonably complete workflow, including dose review and comparison tools
- Require: Labelmap that can handle overlapping structures
- Require: Visualization of RT dose (2D, as isodose lines, with legend)
- Wishlist: Visualization of RT dose (3D, w/ Shadie)
- Require: DICOM improvements (CT export, DICOM-RT import/export)
- Require: Compute dose volume histograms
- Recommend: Margin tools
- Wishlist: Visualize dose volume histograms
- Require: Better support for extensions
- Recommend: MRML node for registration
- Recommend: MRML node for patient demographics
- Wishlist: Unevely spaced CT
- Wishlist: Vector field visualization
- Wishlist: DICOM Network I/O
Data
- Goal: 50 cases for inter-subject segmentation
- Goal: 30 cases for intra-subject registration
- Require: New retrospective IRB
- Require: Careful QA of segmentation data
- Require: Validation data for registration studies
Outreach
- Require: Symposium or tutorial at major conference
- Require: Hands-on tutorial at MGH
- Require: On-line tutorials for radiotherapy workflow
- Recommend: User's group meeting at major conference
Personnel
- MGH: Greg Sharp, Annie Chan, George TY Chen, Nadya Shusharina, Rui Li, Ken Westover, Itai Pashtan, John Wolfgang
- MIT: Polina Golland, Michal Depa
- GT: Allen Tannenbaum, Ivan Kolesov
- Isomics: Steve Pieper