Difference between revisions of "Projects:RegistrationLibrary:RegLib C03"
From NAMIC Wiki
| Line 35: | Line 35: | ||
*[[Image:Button_red_fixed_white.jpg|20px]]reference/fixed : T2w FSE, | *[[Image:Button_red_fixed_white.jpg|20px]]reference/fixed : T2w FSE, | ||
*[[Image:Button_green_moving_white.jpg|20px]] moving: Baseline image of acquired DTI volume, corresponds to T2w MRI , 0.9375 x 0.9375 x 1.4 mm voxel size, oblique | *[[Image:Button_green_moving_white.jpg|20px]] moving: Baseline image of acquired DTI volume, corresponds to T2w MRI , 0.9375 x 0.9375 x 1.4 mm voxel size, oblique | ||
| − | *[[Image: | + | *[[Image:Button_blue_tag_white.jpg|20px]] tag: Tensor data of DTI volume, oblique, same orientation as Baseline image. The result Xform will be applied to this volume. |
=== Registration Results=== | === Registration Results=== | ||
| Line 62: | Line 62: | ||
=== Discussion: Registration Challenges === | === Discussion: Registration Challenges === | ||
| − | * | + | *The DTI contains acquisition-related distortions (commonly EPI acquisitions) that can make automated registration difficult. |
| − | *the two images have strong differences in | + | *the two images often have strong differences in voxel sizes and voxel anisotropy. If the orientation of the highest resolution is not the same in both images, finding a good match can be difficult. |
| − | * | + | *there may be widespread and extensive pathology (e.g stroke, tumor) that might affect the registration if its contrast is different in the baseline and structural reference scan |
| − | |||
=== Discussion: Key Strategies === | === Discussion: Key Strategies === | ||
| − | *the two images have identical contrast, hence we consider "sharper" cost functions, such as NormCorr or MeanSqrd | + | *the two images have identical contrast, hence we could consider "sharper" cost functions, such as NormCorr or MeanSqrd. But because of the strong distortions and lower resolution of the moving image, Mutual Information is recommended as the most robust metric. |
| − | + | *often anatomical labels are available from the reference scan. It would be less work to align the anatomical reference with the DTI, since that would circumvent having to resample the complex tensor data into a new orientation. However the strong distortions are better addressed by registering the other direction, i.e. move the DTI into the anatomical reference space. | |
*because we seek to assess/quantify regional size change, we must use a rigid (6DOF) scheme, i.e. we must exclude scaling. | *because we seek to assess/quantify regional size change, we must use a rigid (6DOF) scheme, i.e. we must exclude scaling. | ||
| − | * | + | *masking is likely necessary to obtain good results. |
| + | *in this example the initial alignment of the two scans is very poor. The strongly oblique orientation of the DTI makes an initial manual alignment step necessary. | ||
*these two images are not too far apart initially, so we reduce the default of expected translational misalignment | *these two images are not too far apart initially, so we reduce the default of expected translational misalignment | ||
| − | *because | + | *because speed is not that critical, we increase the sampling rate from the default 2% to 15%. |
| − | *we also expect | + | *we also expect larger differences in scale & distortion than with regular structural scane: so we significantly (2x-3x) increase the expected values for scale and skew from the defaults. |
| − | + | *a good affine alignment is important before proceeding to non-rigid alignment to further correct for distortions. | |
Revision as of 20:45, 25 January 2010
Home < Projects:RegistrationLibrary:RegLib C03Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
Contents
Slicer Registration Library Exampe #3: Diffusion Weighted Image Volume: align with structural reference MRI
Objective / Background
This is a typical example of DTI processing. Goal is to align the DTI image with a structural scan that provides accuracte anatomical reference. The DTI contains acquisition-related distortion and insufficient contrast to discern anatomical detail.
Keywords
MRI, brain, head, intra-subject, DTI, DWI
Input Data
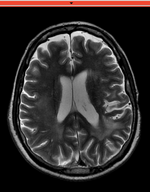
 reference/fixed : T2w FSE,
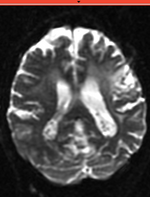
reference/fixed : T2w FSE, moving: Baseline image of acquired DTI volume, corresponds to T2w MRI , 0.9375 x 0.9375 x 1.4 mm voxel size, oblique
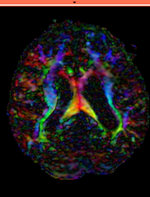
moving: Baseline image of acquired DTI volume, corresponds to T2w MRI , 0.9375 x 0.9375 x 1.4 mm voxel size, oblique tag: Tensor data of DTI volume, oblique, same orientation as Baseline image. The result Xform will be applied to this volume.
tag: Tensor data of DTI volume, oblique, same orientation as Baseline image. The result Xform will be applied to this volume.
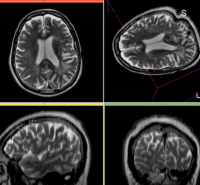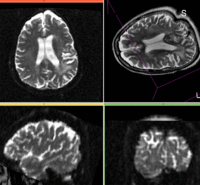
Registration Results
Download
- download entire package (Data,Presets,Tutorial, Solution, zip file 33.7 MB)
- download registration parameter presets file (MRML file, import as scene)
- download image dataset only (NRRD, 10.7 MB, filename: RegLib_C01_Data_TumorGrowth.zip)
- download image dataset only (NRRD, 10.7 MB, filename: RegLib_Case_01_NRRD_TumorGrowth.zip)
- download image dataset in NIFTI format (NIFTI / nii, 10.7 MB, filename: RegLib_C01_DataNIFTI_TumorGrowth.zip)
- result transform file (ITK .tfm file, load into slicer and apply to the target volume)
- Tutorials (step-by -step walk through):
- Multiresolution testresult package (Data,Xform, Solution, zip file 16.5 MB)
- Download package from MIDAS server (Data,Xform)
Discussion: Registration Challenges
- The DTI contains acquisition-related distortions (commonly EPI acquisitions) that can make automated registration difficult.
- the two images often have strong differences in voxel sizes and voxel anisotropy. If the orientation of the highest resolution is not the same in both images, finding a good match can be difficult.
- there may be widespread and extensive pathology (e.g stroke, tumor) that might affect the registration if its contrast is different in the baseline and structural reference scan
Discussion: Key Strategies
- the two images have identical contrast, hence we could consider "sharper" cost functions, such as NormCorr or MeanSqrd. But because of the strong distortions and lower resolution of the moving image, Mutual Information is recommended as the most robust metric.
- often anatomical labels are available from the reference scan. It would be less work to align the anatomical reference with the DTI, since that would circumvent having to resample the complex tensor data into a new orientation. However the strong distortions are better addressed by registering the other direction, i.e. move the DTI into the anatomical reference space.
- because we seek to assess/quantify regional size change, we must use a rigid (6DOF) scheme, i.e. we must exclude scaling.
- masking is likely necessary to obtain good results.
- in this example the initial alignment of the two scans is very poor. The strongly oblique orientation of the DTI makes an initial manual alignment step necessary.
- these two images are not too far apart initially, so we reduce the default of expected translational misalignment
- because speed is not that critical, we increase the sampling rate from the default 2% to 15%.
- we also expect larger differences in scale & distortion than with regular structural scane: so we significantly (2x-3x) increase the expected values for scale and skew from the defaults.
- a good affine alignment is important before proceeding to non-rigid alignment to further correct for distortions.