Difference between revisions of "Projects:RegistrationLibrary:RegLib C03"
From NAMIC Wiki
m (Text replacement - "http://www.slicer.org/slicerWiki/index.php/" to "https://www.slicer.org/wiki/") |
|||
| (50 intermediate revisions by one other user not shown) | |||
| Line 3: | Line 3: | ||
[[Projects:RegistrationDocumentation:UseCaseInventory|Back to Registration Use-case Inventory]] <br> | [[Projects:RegistrationDocumentation:UseCaseInventory|Back to Registration Use-case Inventory]] <br> | ||
| − | ==Slicer Registration Library | + | == <small>updated for '''v4.1'''</small> [[Image:Slicer4_RegLibLogo.png|150px]] <br>Slicer Registration Library Case #3: Diffusion Weighted Image Volume: align with structural reference MRI== |
=== Input === | === Input === | ||
{| style="color:#bbbbbb; " cellpadding="10" cellspacing="0" border="0" | {| style="color:#bbbbbb; " cellpadding="10" cellspacing="0" border="0" | ||
| Line 17: | Line 17: | ||
|} | |} | ||
| − | === | + | ===Objective / Background === |
| − | + | Goal is to align the DTI image with the structural reference T2 scan that provides accuracte anatomical reference. | |
| − | |||
| − | === | + | === Slicer 4.1 Modules Used === |
| − | + | *[https://www.slicer.org/wiki/Documentation/4.1/Modules/BRAINSFit BrainsFit] | |
| + | *[https://www.slicer.org/wiki/Documentation/4.1/Modules/ResampleDTIVolume Resample DTI Volume] | ||
| + | *[https://www.slicer.org/wiki/Documentation/4.1/Modules/DiffusionTensorEstimation Diffusion Tensor Estimation] | ||
| − | === | + | === Alternate Versions === |
| − | + | *this example covers the most basic form of directly registering a DTI + baseline to a T2. There is another (more advanced) version that show how to address additional issues of a strong initial rotation and strong voxel-anisotropy for the raw DWI image acquired. [[Projects:RegistrationLibrary:RegLib_C03B|You will find the advanced version here]]. | |
| + | *[[Projects:RegistrationLibrary:RegLib_C03_v3|for the Slicer 3.6.3 version of this case see here]] | ||
===Download === | ===Download === | ||
| − | + | *Image Data: | |
| − | * | + | **[[Media:RegLib_C03_Data.zip|'''RegLib_C03_Data''': main registration package: register DTI <small> (Data, Transforms, solutions, zip file 115 MB) </small>]] |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | === | + | === Procedure === |
| − | *''' | + | This assumes you have the following: 1) a T2 reference image, 2) a DTI baseline image and 3) the DTI volume (both obtained from the [https://www.slicer.org/wiki/Documentation/4.1/Modules/DiffusionTensorEstimation Diffusion Tensor Estimation module]). |
| − | # | + | *Image Data: |
| − | + | *'''Overview''': | |
| − | # | + | ::#Using [https://www.slicer.org/wiki/Documentation/4.1/Modules/BRAINSFit General Registraion (BRAINS)]''', register DTI_baseline to T2 (affine+nonrigid) w/o masking |
| − | + | :#Resample the DTI with above transform with the [https://www.slicer.org/wiki/Documentation/4.1/Modules/ResampleDTIVolume Resample DTI Volume] module | |
| − | #open Registration : '' | + | #open [https://www.slicer.org/wiki/Modules:BRAINSFit Registration : ''General Registration (BRAINS)''] module |
| − | ## | + | ##''Input Images'': fixed = T2 , moving = DTI_base |
| − | + | ##''Output Settings'': | |
| − | ## | + | ###''Slicer BSpline Transform'' (create new transform, rename to: "Xf1_DTbase-T2_BSpline") |
| − | ### | + | ###''Slicer Linear Transform'' none |
| − | ### | + | ###''Output Image Volume'' (create new volume, rename to: "DTIbaseline_Xf1" |
| − | ### | + | ##''Registration Phases'': select/check ''Rigid'' , ''Rigid+Scale'', ''Affine'', ''BSpline'' |
| − | + | ##''Main Parameters'': | |
| − | + | ###increase ''Number Of Samples'' to 200,000 | |
| − | ## | + | ###set ''B-Spline Grid Size'' to 5,5,5 |
| − | ## | ||
| − | ## | ||
##Leave all other settings at default | ##Leave all other settings at default | ||
| − | ##click: Apply; runtime < 1 min. | + | ##click: ''Apply''; runtime < 1 min. |
| − | + | #Resample DTI | |
| − | + | #Open the [https://www.slicer.org/wiki/Documentation/4.1/Modules/ResampleDTIVolume Resample DTI Volume] module (found under: All Modules) | |
| − | #Open | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
##Input Volume: select DTI | ##Input Volume: select DTI | ||
| − | ##Output Volume: select '' | + | ##Output Volume: select ''create new Diffusion Tensor Volume'',and rename it to ''DTI_Xf1'' |
##Reference Volume: select ''T2'' | ##Reference Volume: select ''T2'' | ||
| − | ##Transform Parameters: select transform " | + | ##Transform Parameters: select transform node "Xf1_DTI-T2_BSpline", for ''Deformation Field'': none ; '''check the ''displacement'' checkbox''' |
| − | |||
##Leave all other settings at defaults | ##Leave all other settings at defaults | ||
| − | ##Click Apply; runtime | + | ##Click Apply; runtime ~ 2 min. |
| − | + | #set ''T2'' as background and new ''DTI_Xf1'' volume as foreground | |
| − | + | #fade between back- and foreground to see DTI overlay onto the T2 image. Note that you can also fade via holding the OPTION+CMD keys (mac) + dragging left mouse. | |
| − | #set '' | ||
| − | # | ||
| − | + | === Registration Results (click to enlarge) === | |
| − | |||
| − | === Registration Results=== | ||
{| style="color:#bbbbbb; background-color:#333333;" cellpadding="10" cellspacing="0" border="0" | {| style="color:#bbbbbb; background-color:#333333;" cellpadding="10" cellspacing="0" border="0" | ||
| − | |[[Image: | + | |[[Image:RegLib_C03_baseline_unregistered.gif|400px|left]] |
| + | |[[Image:RegLib_C03_baseline_registered.gif|400px|left]] | ||
| + | |[[Image:RegLib_C03_DTI_registered.gif|400px|left]] | ||
|- | |- | ||
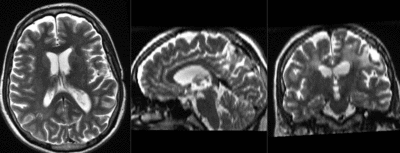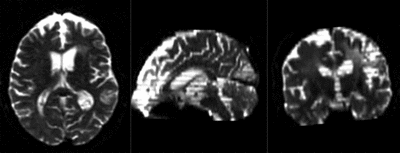
| − | |baseline to T2 after affine alignment | + | |baseline & T2 before registration |
| + | |baseline to T2 after affine+nonrigid alignment | ||
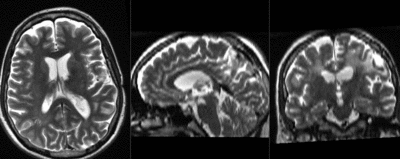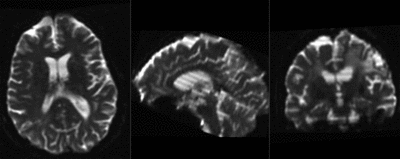
| + | |DTI and T2 before & after registration | ||
|} | |} | ||
| − | + | === Keywords === | |
| − | + | MRI, brain, head, intra-subject, DTI, DWI | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | === | ||
| − | |||
| − | |||
| − | |||
=== Discussion: Key Strategies === | === Discussion: Key Strategies === | ||
| − | * | + | *the strong EPI-based distortions of the DTI image make nonrigid registration necessary |
| − | * | + | *initial alignment & overlap is sufficient so that no "initialization" methods are necessary and registration can succeed without. |
| − | + | *contrast & initial pose are similar enough for registration to succeed without any masking. However the DTI estimation procedure '''does''' provide an optional mask that is usually very helpful in registering cases with more "distracting" image content. [[Projects:RegistrationLibrary:RegLib_C03_2| For an example see the extended version of this case here.]] | |
| − | + | *the DTI in this example is isotropic and hence can be resampled directly. If the DTI contains strong anisotropy of ratios 1:3 or greater, reorienting the DTI can lead to strong artifacts (e.g. in axial direction appear as blue cast in the color orientation view). In that case it is necessary to resample the DWI in the original orientation to an isotropic size before reorienting. It may also be advisable to first reorient the DWI and perform the DTI estimation afterwards. | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
=== Acknowledgments === | === Acknowledgments === | ||
Latest revision as of 17:29, 10 July 2017
Home < Projects:RegistrationLibrary:RegLib C03Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
updated for v4.1 
Slicer Registration Library Case #3: Diffusion Weighted Image Volume: align with structural reference MRI
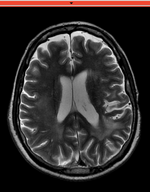
Input
|
|
|
|
| fixed image/target T2 |
moving image 2a DTI baseline |
moving image 2b DTI tensor |
Objective / Background
Goal is to align the DTI image with the structural reference T2 scan that provides accuracte anatomical reference.
Slicer 4.1 Modules Used
Alternate Versions
- this example covers the most basic form of directly registering a DTI + baseline to a T2. There is another (more advanced) version that show how to address additional issues of a strong initial rotation and strong voxel-anisotropy for the raw DWI image acquired. You will find the advanced version here.
- for the Slicer 3.6.3 version of this case see here
Download
- Image Data:
Procedure
This assumes you have the following: 1) a T2 reference image, 2) a DTI baseline image and 3) the DTI volume (both obtained from the Diffusion Tensor Estimation module).
- Image Data:
- Overview:
- Using General Registraion (BRAINS), register DTI_baseline to T2 (affine+nonrigid) w/o masking
- Resample the DTI with above transform with the Resample DTI Volume module
- open Registration : General Registration (BRAINS) module
- Input Images: fixed = T2 , moving = DTI_base
- Output Settings:
- Slicer BSpline Transform (create new transform, rename to: "Xf1_DTbase-T2_BSpline")
- Slicer Linear Transform none
- Output Image Volume (create new volume, rename to: "DTIbaseline_Xf1"
- Registration Phases: select/check Rigid , Rigid+Scale, Affine, BSpline
- Main Parameters:
- increase Number Of Samples to 200,000
- set B-Spline Grid Size to 5,5,5
- Leave all other settings at default
- click: Apply; runtime < 1 min.
- Resample DTI
- Open the Resample DTI Volume module (found under: All Modules)
- Input Volume: select DTI
- Output Volume: select create new Diffusion Tensor Volume,and rename it to DTI_Xf1
- Reference Volume: select T2
- Transform Parameters: select transform node "Xf1_DTI-T2_BSpline", for Deformation Field: none ; check the displacement checkbox
- Leave all other settings at defaults
- Click Apply; runtime ~ 2 min.
- set T2 as background and new DTI_Xf1 volume as foreground
- fade between back- and foreground to see DTI overlay onto the T2 image. Note that you can also fade via holding the OPTION+CMD keys (mac) + dragging left mouse.
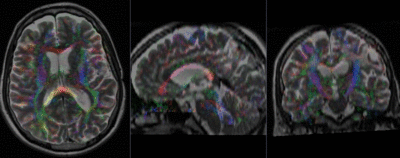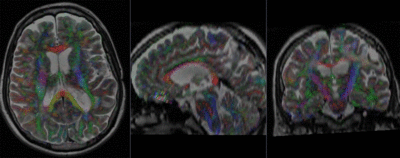
Registration Results (click to enlarge)
| baseline & T2 before registration | baseline to T2 after affine+nonrigid alignment | DTI and T2 before & after registration |
Keywords
MRI, brain, head, intra-subject, DTI, DWI
Discussion: Key Strategies
- the strong EPI-based distortions of the DTI image make nonrigid registration necessary
- initial alignment & overlap is sufficient so that no "initialization" methods are necessary and registration can succeed without.
- contrast & initial pose are similar enough for registration to succeed without any masking. However the DTI estimation procedure does provide an optional mask that is usually very helpful in registering cases with more "distracting" image content. For an example see the extended version of this case here.
- the DTI in this example is isotropic and hence can be resampled directly. If the DTI contains strong anisotropy of ratios 1:3 or greater, reorienting the DTI can lead to strong artifacts (e.g. in axial direction appear as blue cast in the color orientation view). In that case it is necessary to resample the DWI in the original orientation to an isotropic size before reorienting. It may also be advisable to first reorient the DWI and perform the DTI estimation afterwards.