2008 Winter Project Week:GoFigure
 Return to 2008_Winter_Project_Week |
Key Investigators
- Sean Megason (Caltech - Harvard Medical School)
- Alex. Gouaillard (Caltech - Harvard Medical School)
Objective
- Segmentation of cells in 3D+t datasets
- Cross-platform software
Approach, Plan
- leverage NA-MIC Toolkits
- algorithms as ITK classes
- GUI in KW widget
- core to transfer from MFC to std/stl
- keep the access to the DB in the core (avoid MFC::CRecordSet)
Progress
GoFigure
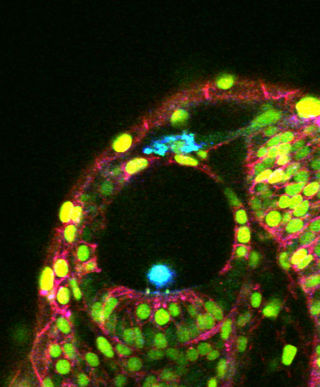
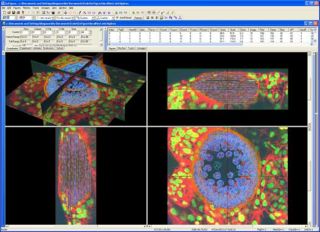
GoFigure is an image analysis software package specifically designed for visualizing, tracking, and analyzing cells in multidimensional confocal/two-photon image sets. Eventually GoFigure will be able to analyze time-lapse series of confocal image stacks (xyzt) and to automatically segment and track cells in 2D, 2D+time, 3D, 3D+time, and 3D+time+cell division.
GoFigure now runs on Microsoft Windows platform. GoFigure utilizes a MySql database back-end so it can handle extremely large image sets.
Open Source and Open Software
the transition phase from the Alpha version (prototype) to a full open source software is almost done. We hope to finish it during the project week. The actual Alpha version is based on MFC and VTK for the vizualization. The Beta version will be based on the following NA-MIC kits: VTK, ITK, CMake(, CTest, CPack) and on MySQL 5.
Progress before project week:
- MySQL 5 prototype ready
- GUI isolated - the KWwidgets we want to use are identified
- design is defined and ready for removing MFC.