2013 Summer Project Week:Sobolev Segmenter
Key Investigators
- UAB: Arie Nakhmani, LiangJia Zhu, Allen Tannenbaum
- BWH: Yi Gao, Ron Kikinis
- Utah: Rob MacLeod, Josh Cates
Objective
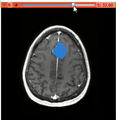
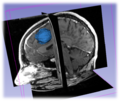
General segmentation tools can be used in a wide range of biomedical applications ranging from tumor delineation to the segmentation of the atrial wall. The latter may be used for the atrial fibrillation DBP. The Sobolev active contour is a smooth general 2D segmenter that can be used in the applications above and is known to be more resistant to noise and local minima those other active contour methodologies. It can be extended for medical volume segmentation. Our objective is to implement Slicer's extension based on Sobolev active contours algorithm for volume segmentation.
Approach, Plan
We prepare C++ implementation of Sobolev active contour algorithm, and convert it to Slicer Commandline extension. This version needs a few sparse initial contours on some slices of the segmented volume.
In another approach, we implement similar algorithm in Python, as an Editor effect of Slicer. In this case, only the definition of 3D depth, and a single click inside the area of the segmented target are needed for the algorithm to work.
Progress
Alpha versions of both approaches (C++ Commandline and Python Editor effect) have been prepared.
Delivery Mechanism
This work will be delivered to the NA-MIC Kit as a loadable Commandline extension and an Editor effect.
References
- A. Nakhmani, A. Tannenbaum, "Tracking with Adaptive Sobolev Snakes." Submitted to IEEE Transactions on Image Processing.
- A. Nakhmani, A. Tannenbaum, "Self-Crossing Detection and Location for Parametric Active Contours," IEEE Transactions on Image Processing, DOI:10.1109/TIP.2012.2188808, Volume 21, Issue 7, pp. 3150-3156, July 2012.