Events: DTI Tractography Challenge MICCAI 2011

| ||

|

|

|
Contents
- 1 DTI Tractography for Neurosurgical Planning: A Grand Challenge
- 2 Overview
- 3 Faculty
- 4 Tentative Agenda
- 5 DTI Tractography Challenge
- 6 Logistics
- 7 Workshop Datasets
- 8 Workshop Format
- 9 How to participate in the Challenge
- 10 Submission Guidelines
- 11 Evaluation
- 12 Suggested background reading
- 13 Post Workshop
DTI Tractography for Neurosurgical Planning: A Grand Challenge
Welcome to the 'DTI Tractography for Neurosurgical Planning: A Grand Challenge' workshop. The goal of this initiative is to provide neurosurgeons with an overview of the tractography methods currently available for reconstructing white matter bundles for pre-surgical planning. The workshop is part of the 14th International Conference on Medical Image Computing and Computer Assisted Intervention MICCAI 2011, to be held from 18th to 22th September 2011 in Toronto, Canada.
Overview
Diffusion Tensor Imaging (DTI) tractography has a unique potential for neurosurgical planning since it provides a window on the complex organization of white matter pathways in-vivo. During the past decade, the MICCAI community has been a major contributor to the development and refinement of a wide variety of advanced tractography techniques. Still the transfer of these cutting-edge algorithms to clinical routine is hindered by the difficulties of validating tractography results. The DTI Tractography Challenge workshop will give participants the opportunity to evaluate the performances of their tractography algorithms in a neurosurgical context. Participants will gain insights on the currently available gold-standard for evaluating tractography results in the Operating Room, in the absence of ground truth.
Faculty
- Sonia Pujol, Ph.D., Surgical Planning Laboratory, Brigham and Women’s Hospital, Harvard Medical School
- Ron Kikinis, M.D., Surgical Planning Laboratory, Brigham and Women’s Hospital, Harvard Medical School
- Alexandra Golby, M.D., Department of Neurosurgery, Brigham and Women’s Hospital, Harvard Medical School
- Guido Gerig, Ph.D.,The Scientific Computing and Imaging Institute, University of Utah
- Martin Styner, Ph.D., Neuro Image Research and Analysis Laboratory, University of North Carolina
- William Wells, Ph.D., Surgical Planning Laboratory, Brigham and Women’s Hospital, Harvard Medical School
- Carl-Fredrik Westin, Ph.D., Laboratory of Mathematics in Imaging, Brigham and Women’s Hospital, Harvard Medical School
- Sylvain Gouttard, M.Sc.,The Scientific Computing and Imaging Institute, University of Utah
- Arya Nabavi, M.D., Department of Neurosurgery, University Hospital Schleswig-Holstein, Kiel, Germany
- Hatsuho Mamata, M.D., Ph.D., Department of Radiology, Brigham and Women’s Hospital, Harvard Medical School
Tentative Agenda
- 8:30-10:00: Start of the competition on two neurosurgical cases: handing out password for data zip file, main processing time
Note: Please follow the submission format guidelines.
- 10:00-10:05: Welcoming remarks and Introduction
- 10:15-10:30: DTI Tractography validation challenge (Sonia Pujol), introduction
- 10:30-10:35: Presentation of the neurosurgical cases and healthy subjects datasets (Alex Golby/Sonia Pujol/ Sylvain Gouttard)
- 10:35-10:45: Coffee-Break
- 10:45-12:15: Tractography Session: Presentation of methods and results on pre-workshop datasets by contestants
- 10:45-10:55: Multifiber Deterministic Streamline Tractography of the Corticospinal Tract Based on a New Diffusion Model. Olivier Commonwick, INRIA Rennes, France
- 10:55-11:05: Involving machine learning and particule mass in the segmentation of cortico-spinal tract. Caroline Brun, UPenn, Philadelphia, USA
- 11:05-11:15: Automated atlas-based seeding in cortico-spinal tractography. Maged Goubran, Robarts Research Institute, Toronto, Canada
- 11:15-11:25: A Volumetric Apprach to extract Corticospinal tract in Diffusion Tensor MRI. Gopal Veni, Scientific Computing and Imaging Institute, Salt Lake City, USA
- 11:25-11:35: Fiber Tracking on Corticospinal Tract in Human Brain using an Intrinsic Unscented Kalman Filter. Guang Cheng, University of Florida, USA
- 11:35-11:45: Global fiber-tracking based on Finsler distance. Antonio Tristan-Vega, Laboratory of Mathematics in Imaging, Boston, USA
- 11:45-11:55: MITK Global Tractography - Application to the Corticospinal Tract. Peter Neher, German Cancer Research Centre, Heidelberg, Germany
- 11:55-12:05: Using Filtered Multitensor Tractography. Yundi Shi, UNC Chapell Hill, USA
- 12:05-13:30: Lunch Break
- 13:30: End of the Grand Challenge on two neurosurgical cases
- 13:30-14:30: Clinical case review: presentation of the Grand Challenge cases by contestants
- 14:30-15:15: Keynote Speaker Lecture: "IG Surgery, evolution, practice and current challenges" Dr. Arya Nabavi, University Hospital Schleswig-Holstein, Kiel, Germany
- 15:15-16:15: Coffee Break and closed-review session
- 16:15-16:40: Presentation of the technical and clinical evaluation criteria (Sylvain Gouttard, Guido Gerig, Martin Styner, Sonia Pujol)
- 16:40-17:00: Presentation of the results of the review
- 17:00-17:30: Announcement of the results and discussion
DTI Tractography Challenge
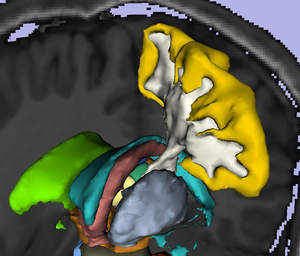
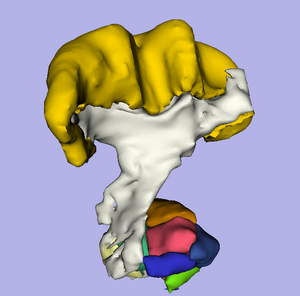
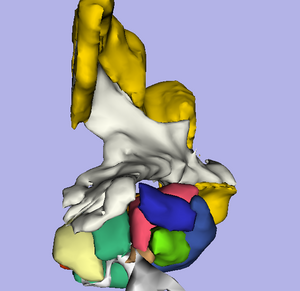
The location and integrity of eloquent white matter pathways is of major importance during neurosurgical planning in order to prevent neurological deficit. Tractography has the potentiel to bring valuable information on tumor infiltration and tract displacement to neurosurgeons. This challenge will focus on the reconstruction of the major pathway for voluntary motor function, the corticospinal tract (click here for definition). Participants will be required to reconstruct the left and right corticospinal tracts in datasets acquired on four neurosurgical cases and two control subjects. Participants will be invited to submit their tractography results for two neurosurgical datasets and two control subjects before the workshop, and to process two neurosurgical cases at the event. Both positive and negative tractography findings will be assessed using qualitative and quantitative criteria. In addition, the tracking reliability will be assessed on the repeated volunteer scans.
Logistics
The DTI Tractography Challenge workshop will be held on Sunday September 18, 2011 at the Westin Harbour Castle, 1 Harbour Square, Toronto, Canada.
Registration: Space is limited. To register for the workshop, please visit the MICCAI website.
Workshop Datasets
The workshop datasets consist of neurosurgical cases provided by the Department of Neurosurgery at Brigham and Women's Hospital, Boston, MA, and the repeated volunteer scans from two healthy subjects provided by The Scientific Computing and Imaging Institute, Salt Lake City, Utah. Each dataset includes Anatomical Images (T1,T2), a Diffusion Weighted Imaging (DWI) volume, and a Diffusion Tensor Imaging (DTI) volume. Each clinical dataset includes a segmentation of the tumor. Data are in the ITK-readable Nrrd file format, which consists of an ASCII header file and a separate uncompressed raw image datafile.
Workshop Format
The workshop will be composed of two parts: the first part will consist of a series of presentations of the tractography algorithms and results by the workshop participants, in parallel with challenge contest; the second part will focus on the results of the contest, along with a series of lectures and discussions on tractography challenges and clinical use of DTI for neurosurgical planning.
How to participate in the Challenge
- Update: The workshop datasets are available.
- To participate to the DTI Tractography Challenge, download and fill the Letter of Intent for Submission, and send it by email to Sonia Pujol (spujol at bwh.harvard.edu). After successful registration of your team, we will send you the link to the workshop data.
Submission Guidelines
- Participants will be required to submit a zip archive file containing their tractography results for the neurosurgical cases and for the healthy subjects, along with a short paper. Please visit the submission format page for the file format details.
- The tractography results should include 1) the 3D coordinate of the tracts in the VTK-ASCII file format (vtkPolyData), 2) the envelope of the tracts in the ITK-readable Nrrd file format, 3) the regions of interest used for the reconstruction (if applicable) in the ITK-readable Nrrd file format, and 4) three anatomical views (axial, coronal, sagittal) of the tract in png file format.
- The paper should include the following elements 1) a presentation of the DTI analysis pipeline, 2) a description of the tractography techniques and parameters used, 3) a set of images providing an intuitive way to present the reconstructed tracts to neurosurgeons. Participants will be invited to propose an evaluation criteria for their tractography method in the Appendix of the paper.
- Supplementary materials such as short videos, are encouraged, but not mandatory.
- Papers should be formatted in Lecture Notes in Computer Science style. The file format for submission is Adobe Portable Document Format (PDF). Other formats will not be accepted.
- Once you have completed the analysis, upload your workshop paper and results at the location of your choice, and send the url to access them to Sonia Pujol. Within 48 hours, you'll receive a confirmation of your submission.
Important Dates:
Dead-line for submission: July 1st, 2011 (extended)
Notification of Acceptance: July 11, 2011
Camera-ready papers: August 1, 2011
Evaluation
Tractography results will be evaluated based on a set of qualitative and quantitative criteria. Qualitative evaluation will be performed by three experts.
- The qualitative assessment of tract reconstruction in each hemisphere, for patients and controls, will be based on 1) anatomical correctness of the tract, 2) presence of false-positive tracts, 3) presence of false-negative tracts.
- The qualitative assessment of the depiction of the spatial relation between the tumor and the tract will be based on: 1) correct depiction of the distance between the tract and the lesion, 2) demonstration of tract displacement, and 3) demonstration of tumor infiltration.
- The quantitative evaluation of tract reconstruction in each hemisphere, for patients and controls, will be performed using five metrics : 1) Dice's coefficient for volumetric overlap, 2) Closest point distance between bundles (Hausdorff distance), 3) Fiber profiles on Fractional Anisotropy (FA), 4) Fiber profiles of Mean Diffusivity (MD), 5) STAPLE sensitivity and specificity score.
Suggested background reading
- MRI Atlas of Human White Matter. S. Mori, S. Wakana, P.C.M. van Zijl, L.M. Nagae-Poetscher. Elsevier Science, 2005
- Diffusion MRI: Theory, Methods, and Applications. Edited by Derek K. Jones. Oxford University Press, 2011.
Post Workshop
For the on-going work and outcomes of the MICCAI 2011 workshop, please see the post-workshop page and workshop proceedings
- Cortico Spinal Tract (CST), from the 2008 edition of the PNL SPL brain atlas
- The atlas is freely available. For a download, see here. In addition to the label maps, it is configured to be loaded into 3D Slicer. Several preconfigured views are available.
- For more information about where the CST is located in the posterior limb in the internal capsule and spinal cord, have a look at the picture in this link.