Projects:RegistrationLibrary:RegLib C38
From NAMIC Wiki
Home < Projects:RegistrationLibrary:RegLib C38
updated for v4.1
Back to ARRA main page
Back to Registration main page
Back to Registration Use-case Inventory
Contents
updated for v4.1  Slicer Registration Library Case 38: TBI
Slicer Registration Library Case 38: TBI
Input
|
|
|
|
|
|
|
|
|
|
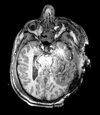
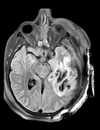
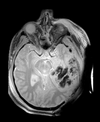
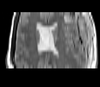
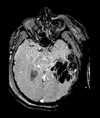
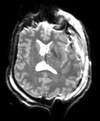
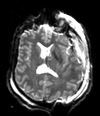
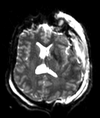
| T1pre | FLAIR | T1post | coronal T2 | SWI | EP50 | EP75 | EP100 |
Slicer 4.1 recommended Modules
Download
- TBI case: (Data & Solution Xforms, zip file 40 MB)
- Documentation/Tutorials: short video clips (no sound)
- register T2-T1: Media:RegLib_C38_T2_register.mov (quicktime movie, 17 MB)
- register T2-SWI: Media:RegLib_C38_SWI_register.mov (quicktime movie, 14 MB)
- register T2-GRE: Media:RegLib_C38_GRE_register.mov (quicktime movie, 14 MB)
- register T2-FLAIR: Media:RegLib_C38_FLAIR_register.mov (quicktime movie, 9 MB)
- register T2-EP: Media:RegLib_C38_EP_register.mov (quicktime movie, 9 MB)
- apply all Xforms: Media:RegLib_C38_ApplyXform.mov (quicktime movie, 6MB)
Objective / Background
We seek to align two exams (acute baseline and follow-up) as well as all series within exams into a common space for direct evaluation of regional change.
Input Data
- inter-exam
- reference/fixed : T1_e1 baseline exam of acute TBI , 512 x 512 x 160 ; 0.5 x 0.5 x 1 mm voxel size, axial acquisition, RAS orientation.
- moving:T1_e2 follow-up exam of acute TBI , 512 x 512 x 160 ; 0.5 x 0.5 x 1 mm voxel size, axial acquisition, RAS orientation.
- intra-exam
Registration Challenges
- extensive injury
- distortions on EPI images
- clipped FOV on T2
Key Strategies
- we register all the scans within each exam to the T1
- we limit registration degree of freedom to 6 DOF (rigid transformation only).
Procedure
- Load all datasets via drag&drop or via "AddData / AddVolume" ... do not center the volumes, since they have reasonable alignment at the outset
- the registration procedure below is identical for all other scans in the sequence, i.e. just replace the Moving Volume and the Slicer Linear Transform", everything else can stay at default
- open the General Registration (BRAINS) module
- Fixed Image Volume: T1
- Moving Image Volume: T2
- Output Settings:
- Slicer BSpline Transform": none
- Slicer Linear Transform: create new transform, rename to "Xf1_T2"
- Output Image Volume: none
- Registration Phases: check box for Rigid only
- leave all other settings at defaults
- click: Apply; runtime < 15 sec (MacPro QuadCore 2.4GHz)
- this will generate the alignment transform. Note that the Slicer4.1 version of BRAINS registration does not automatically place the moving volume inside the result transform, as it did for Slicer 3.6. In order to see the result, you must either go to the Data module and drag the moving volume inside the transform node, or select to generate a resampled Output Volume, as done above. The demo movies you can download here show how to do this.

