Projects:RegistrationTBI
Back to Stony Brook University Algorithms
Multimodal Deformable Registration of Traumatic Brain Injury MR Volumes using Graphics Processing Units
Description
An estimated 1.7 million Americans sustain traumatic brain injuries (TBI's) every year. The large number of recent TBI cases in soldiers returning from military conflicts has highlighted the critical need for improvement of TBI care and treatment, and has drawn sustained attention to the need for improved methodologies of TBI neuroimaging data analysis. Neuroimaging of TBI is vital for surgical planning by providing important information for anatomic localization and surgical navigation, as well as for monitoring patient case evolution over time. Approximately 2 days after the acute injury, magnetic resonance imaging (MRI) becomes preferable to computed tomography (CT) for the purpose of lesion characterization, and the use of various MR sequences tailored to capture distinct aspects of TBI pathology provides clinicians with essential complementary information for the assessment of TBI-related anatomical insults and pathophysiology.
Image registration plays an essential role in a wide variety of TBI data analysis workflows. It aims to find a transformation between two image sets such that the transformed image becomes similar to the target image according to some chosen metric or criterion. Typically, a similarity measure is first established to quantify how `close` two image volumes are to each other. Next, the transformation that maximizes this similarity is typically computed through an optimization process which constrains the transformation to a predetermined class, such as rigid, affine or deformable. Numerous challenges associated with the task of TBI volume co-registration can exist if data acquisition is performed multimodally, and additional complexities can also arise due to the large degree of algorithmic robustness that may be required in order to properly address pathology-related deformations. Many conventional methods use the sum of squared differences of intensity values between two image sets as a similarity measure, which can perform poorly or even fail for TBI volume registration. Consequently, because the deformation of patient anatomy and soft tissues cannot typically be represented by rigid transforms, image registration often requires deformable image registration (DIR), i.e., the necessity of applying nonparametric infinite-dimensional transformations.
This paper proposes to replace the Mutual Information (MI) criterion for registration with the Bhattachayya distance [1] within a multimodal DIR framework [3]. The advantage of BD over MI is the superior behavior of the square root function compared to that of the logarithm at zero, which yields a more stable algorithm. This framework we describe takes into account the physical models of tissue motion to regularize the deformation fields and also involves free-form deformation. On the other hand, the DIR algorithm is computationally expensive when implemented on conventional central processing units, which can be detrimental particularly when three-dimensional (3D) volumes-rather than 2D images-need to be co-registered. In clinical settings that involve acute TBI care, the amount of time required by the processing of neuroimaging data sets from patients in critical condition should be minimized. To meet this clinical requirement, we have implemented our algorithm on a graphics processing unit (GPU) platform [2].
Result
MR volumes were acquired at 3 T using a Siemens Trio TIM scanner (Siemens AG, Erlangen, Germany). Because assessing the time evolution of TBI between the acute to the chronic stage is of great interest in the clinical field in order to evaluate case evolution, scanning sessions were held both several days (acute baseline) as well as 6 months (chronic follow-up) after the traumatic injury event. To eliminate the effect of different scanner parameters during each scanning session, every subject was scanned using the same scanner for both acute and chronic time points. The MP-RAGE sequence (Mugler and Brookeman, 1990) was used to acquire T1-weighted images. In addition, MR data were also acquired using fluid-attenuated inversion recovery (FLAIR, (De Coene et al., 1992)), gradient-recalled echo (GRE) T2-weighted images as well as diffusion weighted imaging (DWI), and perfusion imaging.
Before applying our deformable registration algorithm, all image volumes were co-registered by rigid-body transformation to the pre-contrast T1-weighted volume acquired during the acute baseline scanning session. This helps to correct for head tilt and reduce error in computing the local deformation fields. Another technique that was employed before performing the registration is skull stripping, which was useful in our case because images acquired at the acute stage exhibit appreciably more extracranial swelling compared to images acquired chronically. Since all modalities are co-registered to T1, we only need to perform the skull stripping once, i.e. on the T1 volume. Skull stripping is necessary because, without it, the DIR algorithm would deform the interior of the brain to match the outside boundary. This type of deformation is mathematically valid, but does not yield anatomically plausible results. Two possible solutions to this problem are either adding prior knowledge on the boundary or applying skull stripping, of which we opt for the latter due to its common usage in image processing. We use the BrainSuit software [1] for the skull stripping.
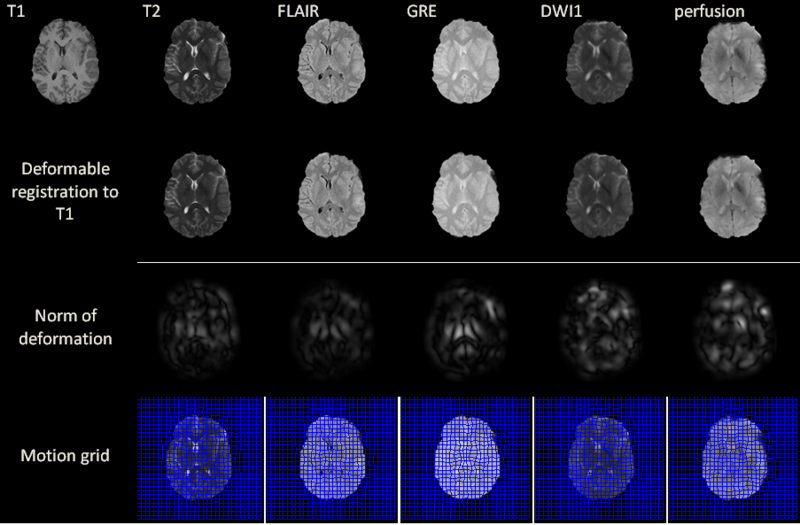
Excluding preprocessing steps, the registration of two volumes of sizes 256x256x60 is found to require 6 seconds on the GPU. Registration results are illustrated for a 2D slice in the above figure for acute stage. The norm of the deformation fields and its 2D motion grid are also included. For T2, FLAIR and GRE volumes, the largest amount of deformation is observed bilaterally in the deep periventricular white matter, possibly as a consequence of hemorrhage and/or CSF infiltration into edemic regions, which can alter voxel intensities in GRE and FLAIR imaging, respectively. In the case of DWI, notable deformation is observed frontally and frontolaterally; in the former case, this may be the result of warping artifacts due to the large drop in the physical properties of tissues at the interfaces between brain, bone and air. In the latter case, the deformation is possibly due to the presence of TBI-related edema, which can substantially alter local diffusivity values. Similar effects due to these causes are observed with DWI and with perfusion imaging in both acute and chronic scans.
Future Work
Future work will focus on registration of TBI volumes across time in terms of registering acute to chronic or vice versa. There are a large number of registration algorithms that assume the smoothness of the vector flow, i.e., the deformation is diffeomorphic. However, when registering TBI across time, the deformation is not well-defined, let alone to be diffeomorphic, at some regions where bleeding or lesion occurs. It is challenging and important to design a registration algorithm that can deal with topological changes for TBI patients. One possible approach is to use combination of locally rigid and non-rigid transforms based on visual features such as MIND descriptor [4]. Some regions of new lesions cannot be explained by minor elastic registration, thus simultaneous registration and segmentation of lesions is needed. We are working on the improved volume matching with boundary conditions based on this segmentation.
Key Investigators
Georgia Tech: Yifei Lou and Patricio Vela
UAB: Arie Nakhmani and Allen Tannenbaum
UCLA: Andrei Irimia, Micah C. Chambers, Jack Van Horn and Paul M. Vespa
References
1.Yifei Lou, Andrei Irimia, Patricio Vela, Micah C. Chambers, Jack Van Horn, Paul M. Vespa and Allen Tannenbaum. Multimodal Deformable Registration of Traumatic Brain Injury MR Volumes via the Bhattacharyya Distance. Submitted to IEEE Transactions on Bioengineering, 2012
2. Yifei Lou, Xun Jia, Xuejun Gu and Allen Tannenbaum. A GPU-based Implementation of Multimodal Deformable Image Registration Based on Mutual Information or Bhattacharyya Distance. Insight Journal, 2011. [2]
3. E. D’Agostino, F. Maes, D. Vandermeulen, and P. Suetens. A viscous fluid model for multimodal non-rigid image registration using mutual information,” MICCAI, 2002, pp. 541–548
4. M.P. Heinrich, M. Jenkinson, M. Bhushan, T. Matin, F. Gleeson, M. Brady, J.A. Schnabel. MIND: Modality independent neighbourhood descriptor for multi-modal deformable registration. Medical Image Analysis. Vol. 16(7) Oct. 2012, pp. 1423–1435, Special Issue on MICCAI 2011