2013 Project Week:WebbasedAnatomicalTeachingFrameworkSummer2013
From NAMIC Wiki
Home < 2013 Project Week:WebbasedAnatomicalTeachingFrameworkSummer2013
Key Investigators
- Nathaniel Reynolds, Randy Gollub, Lilla Zollei (MGH)
- Rudolph Pienaar, Ellen Grant, Daniel Haehn, Nicolas Rannou (Boston Children's Hospital)
- Steve Pieper (Isomics Inc.)
Project Description
Objective
- Over the last several years, an infrastructure has been developed to integrate health records and PACS image archives to support research on large collections of data collected during delivery of healthcare (as contrasted with data collected as part of clinical trials). This resulted in the availability of the mi2b2 system. An application enabled by this technology is supported by an R01 to use clinical scans of children aged zero to six years old to build a teaching atlas.
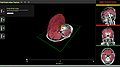
- One goal of the mi2b2 collaborative effort is to create a web-based teaching tool for fetal and neonatal anatomy as well as a browser for quantified information. A basic prototype has been created: http://fnndsc.github.io/babybrain/. The purpose of this project is to construct a 4D MRI dataset as a concatenation of multiple 3D MRI datasets (pediatric MRI scans at different stages of development), and to develop the functionality within XTK to visualize and navigate this 4D dataset in a web browser.
Approach, Plan
Right now, the multiple volumes of the training data get loaded one-by-one on demand. We want to load either everything at the beginning (maybe from a 4D dataset or separate 3D ones) or leverage webworkers to load the rest odf the data in the background while the user interacts with the page.
Progress
We leveraged Web Workers and the new developments of the XIO to download multiple MRI datasets in parallel and not interfere with operation of the main thread. An example can be found here: http://sandbox.goxtk.com/xio/ Our next move is to incorporate these developments into our Pedi-Brain Atlas Learning Prototype.
Draft Development Plans
Phase I: Developer Mockups
During the first half of 2013:
- ...
Phase II: Limited Beta
During second half of 2013:
- ...
Phase III: Authoring Tools
During 2014:
- ...