Difference between revisions of "Algorithm:UNC"
| (81 intermediate revisions by 6 users not shown) | |||
| Line 3: | Line 3: | ||
= Overview of UNC Algorithms (PI: Martin Styner) = | = Overview of UNC Algorithms (PI: Martin Styner) = | ||
| − | At UNC, we are interested in a range of algorithms and solutions for the surface based analysis of brain structures and the cortex. We pioneered the use of spherical harmonics based shape analysis for comparing brain structures across objects. We | + | At UNC, we are interested in a range of algorithms and solutions for the surface based analysis of brain structures and the cortex. We pioneered the use of spherical harmonics based shape analysis for comparing brain structures across objects. We has also worked on incorporating various data sources for correspondence computation on surfaces of different complexity (ranging from simple brain structures to the highly folded cortical surface). A current topic includes the use of diffusion tensor imaging for connectivity analysis in pathological settings. Finally, investigating quality control, validation and evaluation methodology is another important topic of our NA-MIC research. |
= UNC Projects = | = UNC Projects = | ||
| Line 9: | Line 9: | ||
{| cellpadding="10" style="text-align:left;" | {| cellpadding="10" style="text-align:left;" | ||
| + | |||
| + | | style="width:15%" |[[Image:UNC_DTIAnalysisFramework.jpg|200px]] | ||
| + | | style="width:85%" | | ||
| + | |||
| + | == [[Projects:AtlasBasedDTIFiberAnalyzerFramework| Atlas Based DTI Fiber Analysis Framework]] == | ||
| + | This project aims to define an automatic framework for statistical comparison of fiber bundle diffusion properties between populations of diffusion weighted images. | ||
| + | |||
| + | Quality control is performed on diffusion weighted images and DTI images are computed for each individual subject. The data is then either mapped into a prior atlas or an unbiased diffeomorphic DTI atlas is generated from all datasets. This creates a normalized coordinate system for all diffusion images in a study. Fiber tracts of interest are generated on this atlas, and then mapped back to the individual subjects. Diffusion properties along fiber tracts, such as fractional anisotropy (FA), are modeled as multivariate functions of arc length and gathered in spreadsheets for statistical analysis. | ||
| + | [[Projects:AtlasBasedDTIFiberAnalyzerFramework|More...]] | ||
| + | |||
| + | <font color="red">'''New: '''</font> Update versions on NITRC for [http://www.nitrc.org/projects/dti_tract_stat/ DTIAtlasFiberAnalyzer] and [http://www.nitrc.org/projects/fvlight/ FiberViewerLight ] and [http://www.nitrc.org/projects/niral_utilities DTIAtlasBuilder] | ||
| + | |||
| + | <font color="red">'''New: '''</font> AR Verde, JB Berger, A Gupta, M Farzinfar, A Kaiser, VW Chanon, C Boettiger, C Goodlett, Y Shi, H Zhu, G Gerig, S Gouttard, C Vachet, M Styner, The UNC-Utah NA-MIC DTI framework: Atlas Based Fiber Tract Analysis with Application to a Study of Nicotine Smoking Addiction, accepted to SPIE Medical Imaging 2013 | ||
| + | |||
| + | |- | ||
| + | |||
| + | | style="width:15%" |[[Image:UNC_longitudinalAtlasEx1.png|200px]] | ||
| + | | style="width:85%" | | ||
| + | |||
| + | == [[Projects:LongitudinalAtlasBuilding| Longitudinal Atlas Building]] == | ||
| + | As part of the longitudinal intra- and interpatient analysis theme within NA-MIC, we are working on a deformable, longitudinal DTI atlas method. Our longitudinal framework explicitly accounts for temporal dependencies via iterative subject-specific statistical growth modeling, and cross-sectional atlas-building. To effectively account for measurements sparse in time, a continuous-discrete statistical growth model is proposed incorporating also patient co-variates[[Projects:LongitudinalAtlasBuilding|More...]] | ||
| + | |||
| + | <font color="red">'''New: '''</font> I. Csapo, B. Davis, Y. Shi, M. Sanchez, M. Styner, and M. Niethammer, Temporally-dependent image similarity measure for longitudinal analysis, presented at the WBIR'12: Proceedings of the 5th international conference on Biomedical Image Registration, 2012, vol. 7359, pp. 99–109. | ||
| + | |||
| + | <font color="red">'''New: '''</font> I. Csapo, B. Davis, Y. Shi, M. Sanchez, M. Styner, and M. Niethammer, “Longitudinal Image Registration with Non-uniform Appearance Change,” Medical Image Computing and Computer-Assisted Intervention – MICCAI 2012, vol. 7512, pp. 280–288, Aug. 2012. | ||
| + | |||
| + | <font color="red">'''New: '''</font> Y. Hong, S. Joshi, M. Sanchez, M. Styner, and M. Niethammer, Metamorphic Geodesic Regression, Medical Image Computing and Computer Assisted Interventions MICCAI 2012, vol. 7512, pp. 197–205, Aug. 2012. | ||
| + | |||
| + | |- | ||
| + | |||
| + | | style="width:15%" |[[Image:UNC_dwiatlas.png|200px]] | ||
| + | | style="width:85%" | | ||
| + | |||
| + | == [[Projects:DWIAtlas|Diffusion Weighted Atlas Construction via model-based transformation and averaging of signal]] == | ||
| + | This project investigated a method for model-based averaging of sets of diffusion weighted magnetic resonance images (DW-MRI) under space transformations (resulting for example from registration methods). A robust weighted least squares method is developed. Synthetic validation experiments show the improvement of the proposed estimation method in comparison to standard least squares estimation. The developed method is applied to construct an atlas of {\it diffusion weighted images} for a set of macaques, allowing for a more flexible representation of average diffusion information compared to standard diffusion tensor atlases. | ||
| + | [[Projects:DWIAtlas|More...]] | ||
| + | |||
| + | |||
| + | <font color="red">'''New: '''</font> Y. Shi, S. Benzaid, M. Sanchez, M. Styner, M. Niethammer. Diffusion Weighted Atlas Construction via robust model-based transformation. NeuroImage, submitted. | ||
| + | |||
| + | <font color="red">'''New: '''</font> M. Niethammer, Y. Shi, S. Benzaid, M. Sanchez, and M. Styner. Robust model-based transformation and averaging of diffusion weighted images applied to diffusion weighted atlas construction. MICCAI, Workshop on Computational Diffusion MRI, 2010. | ||
| + | |||
| + | |- | ||
| + | |||
| + | | style="width:15%" |[[Image:UNC_GraphbasedConnectivity_Ex1.png|200px]] | ||
| + | | style="width:85%" | | ||
| + | |||
| + | == [[Projects:DiffusionGraphBasedConnectivity|Diffusion Imaging based Connectivity]] == | ||
| + | |||
| + | This project focuses on connectivity measurements derived from diffusion imaging datasets in order to better understand cortical and subcortical white matter connectivity. Our research employs a novel, multi-directional graph propagation method that performs a fully deterministic, efficient and stable connectivity computation. The method handles crossing fibers and deals well with multiple seed regions. In addition to the analysis of these connectivity measures in describing brain pathology, they can also be used as scalar maps for use in DTI registration. | ||
| + | [[Projects:DiffusionGraphBasedConnectivity|More...]] | ||
| + | |||
| + | <font color="red">'''New: '''</font> I. Oguz, A. Boucharin, W. Lu, C. Vachet, F. Budin, Y. Shi, and M. Styner, A Minimum Cost Approach to Connectivity from Orientation Distribution Functions via Efficient Multi-directional Graph Propagation, Proceedings of CDMRI workshop, MICCAI 2012, pp. 210–221, Dec. 2012. | ||
| + | |||
| + | <font color="red">'''New: '''</font> Alexis Boucharin, Ipek Oguz, Clement Vachet, Yundi Shi, Mar Sanchez, Martin Styner. Efficient, graph-based white matter connectivity from orientation distribution functions via multi-directional graph propagation. Medical Imaging 2011: Image Processing (2011) vol. 7962 (1) pp. 79620S | ||
| + | |||
| + | |- | ||
| + | |||
| + | | style="width:15%" |[[Image:DTIPrep_example1.png|200px]] | ||
| + | | style="width:85%" | | ||
| + | |||
| + | == [[Projects:DTI_DWI_QualityControl|DWI and DTI Quality Control]] == | ||
| + | |||
| + | DWI data suffers from inherent low SNR, overall long scanning time of multiple directional encoding with correspondingly large risk to encounter several kinds of artifacts. These artifacts can be too severe for a correct and stable estimation of the diffusion tensor. Thus, a quality control (QC) procedure is absolutely necessary for DTI studies. We are developing a framework for automatic DWI and DTI quality assessment and correction. We developed a tool called DTIPrep which pipelines the QC steps with designated protocol use and report generation. [[Projects:DTI_DWI_QualityControl|More...]] | ||
| + | |||
| + | <font color="red">'''New: '''</font> Mahshid Farzinfar, Yinpeng Li, Audrey Verde, Ipek Oguz, Guido Gerig, Martin A. Styner, DTI quality control assessment via error estimation from Monte Carlo simulation, to be presented at SPIE Medical Imaging 2013. | ||
| + | |||
| + | <font color="red">'''New: '''</font> M. Farzinfar, C. Dietrich, R. G. Smith, Y. Li, A. Gupta, Z. Liu, and M. Styner, “Entropy based DTI quality control via regional orientation distribution,” Biomedical Imaging (ISBI), 2012 9th IEEE International Symposium on, pp. 22–25, 2012. | ||
| + | |||
| + | <font color="red">'''New: '''</font> DTIPrep new version on [http://www.nitrc.org/projects/dtiprep/ NITRC ], including visual quality control support | ||
| + | |||
| + | |- | ||
| + | |||
| + | | style="width:15%" |[[Image:PrimateSulcalCorrUNC.jpg|200px]] | ||
| + | | style="width:85%" | | ||
| + | |||
| + | == [[Projects:CorticalCorrespondenceWithSulcalCurves|Cortical Correspondence using Sulcal Curves]] == | ||
| + | |||
| + | In this project, we compute group wise cortical correspondence on populations, using information from sulcal curves, as well as a novel way to encode deformations on the sphere. This framework is then applied to data from human and non-human primate cortical thickness studies. [[Projects:CorticalCorrespondenceWithSulcalCurves|More...]] | ||
| + | |||
| + | <font color="red">'''New: '''</font>Ilwoo Lyu, Sun Hyung Kim, Joon-Kyung Seong, Soongsil Univ., Sang Wook Yoo, Alan Evans, Yundi Shi, Mar Sanchez, Marc Niethammer, Martin A. Styner, Cortical correspondence via sulcal curve-constrained spherical registration with application to Macaque studies, to be presented at SPIE Medical Imaging 2013 | ||
| + | |||
| + | |- | ||
| + | |||
| style="width:15%" |[[Image:Sulcaldepth.png|200px]] | | style="width:15%" |[[Image:Sulcaldepth.png|200px]] | ||
| Line 15: | Line 99: | ||
== [[Projects:CorticalCorrespondenceWithParticleSystem|Cortical Correspondence using Particle System]] == | == [[Projects:CorticalCorrespondenceWithParticleSystem|Cortical Correspondence using Particle System]] == | ||
| − | In this project, we want to compute cortical correspondence on populations, using various features such as cortical structure, DTI connectivity, vascular structure, and functional data (fMRI). This presents a challenge because of the highly convoluted surface of the cortex, as well as because of the different properties of the data features we want to incorporate together. [[Projects:CorticalCorrespondenceWithParticleSystem|More...]] | + | In this project, we want to compute cortical correspondence on populations, using various features such as cortical structure, DTI connectivity, vascular structure, and functional data (fMRI). This presents a challenge because of the highly convoluted surface of the cortex, as well as because of the different properties of the data features we want to incorporate together. This correspondence method has been included in our NAMIC cortical thickness framework GAMBIT [[Projects:CorticalCorrespondenceWithParticleSystem|More...]] |
| − | <font color="red">'''New: '''</font> | + | <font color="red">'''New: '''</font> Vachet, C., Hazlett, H., Niethammer, M., Oguz, I., Cates, J., Whitaker, R., Piven, J., Styner, M., “Group-wise automatic mesh-based analysis of cortical thickness“. Medical Imaging 2011: Image Processing (2011) vol. 7962 (1) pp. 796227 1 - 10 |
| + | |||
| + | <font color="red">'''New: '''</font> Lee J, Ehlers C, Crews F, Niethammer M, Budin F, Paniagua B, Sulik K, Johns J, Styner M, Oguz I. Automatic cortical thickness analysis on rodent brain. Medical Imaging 2011: Image Processing (2011) vol. 7962 (1) pp. 796248 | ||
|- | |- | ||
| − | |||
| Line 26: | Line 111: | ||
| | | | | | ||
| − | == [[Projects:ShapeAnalysisFrameworkUsingSPHARMPDM|Shape Analysis Framework | + | == [[Projects:ShapeAnalysisFrameworkUsingSPHARMPDM|UNC-Utah Shape Analysis Framework]] == |
| − | The UNC shape analysis is based on an analysis framework of objects with spherical topology, described mainly by sampled spherical harmonics SPHARM-PDM. The input of the shape analysis framework is a set of binary segmentations of a single brain structure, such as the hippocampus or caudate. These segmentations are converted into a shape description (SPHARM) with correspondence and | + | The UNC shape analysis is based on an analysis framework of objects with spherical topology, described mainly by sampled spherical harmonics SPHARM-PDM. The input of the shape analysis framework is a set of binary segmentations of a single brain structure, such as the hippocampus or caudate. These segmentations are converted into a shape description (SPHARM-PDM) with correspondence and tested via statistical point-wise analysis. Additionally, the SPHARM correspondences can be improved with Entropy-based particle systems, by using an integration module recently added to the pipeline. [[Projects:ShapeAnalysisFrameworkUsingSPHARMPDM|More...]] |
| − | + | * SPHARM-Particle Shape Analysis Toolkit disseminated on [http://www.nitrc.org/projects/spharm-pdm NITRC SPHARM PDM page]. All tools are Slicer compatible. Slicer extension module in prep. | |
| − | + | <font color="red">'''New: '''</font> N. Solowij, M. Walterfang, D. I. Lubman, S. Whittle, V. Lorenzetti, M. Styner, D. Velakoulis, C. Pantelis, and M. Yücel, “Alteration to hippocampal shape in cannabis users with and without schizophrenia.,” Schizophr Res, Nov. 2012. | |
| − | |||
| − | |||
| − | |||
| − | + | <font color="red">'''New: '''</font> O. Lindberg, M. Walterfang, J. C. L. Looi, N. Malykhin, P. Ostberg, B. Zandbelt, M. Styner, B. Paniagua, D. Velakoulis, E. Orndahl, and L.-O. Wahlund, “Hippocampal Shape Analysis in Alzheimer's Disease and Frontotemporal Lobar Degeneration Subtypes.,” J Alzheimers Dis, Mar. 2012. | |
| − | + | <font color="red">'''New: '''</font> D. Ong, M. A. Walterfang, G. Malhi, M. Styner, D. Velakoulis, and C. Pantelis, “Shape Alteration in the Caudate Nucleus in Individuals With Bipolar Affective Disorder Disorder.,” Aust N Z J Psychiatry, Feb. 2012. | |
| − | |||
| − | = | + | <font color="red">'''New: '''</font> Beatriz Paniagua, Omri Emodi, Jonathan Hill, James Fishbaugh, Luiz Pimenta, Stephen Aylward, Andinet Enquobahrie, Guido Gerig, John Gilmore, John van Aalst, Martin A. Styner, 3D of brain shape and volume after cranial vault remodeling surgery for craniosynostosis correction in infants, to be published in SPIE Medical Imaging 2013 |
| − | + | <font color="red">'''New: '''</font> Beatriz Paniagua, Amanda Lyall, Jean-Baptiste Berger, Clement Vachet, Robert Hamer, Sandra Woolson, Weili Lin, John Gilmore, Martin A. Styner, Lateral ventricle morphology analysis via mean latitude axis, to be published in SPIE Medical Imaging 2013 | |
| − | <font color="red">'''New: '''</font> Styner | + | <font color="red">'''New: '''</font> Eric Maltbie, Kshamta Bhatt, Beatriz Paniagua, Rachel Smith, Michael Graves, Matthew Mosconi, Sarah Peterson, Scott White, Joseph Blocher, Mohammed El-Sayed, Heather Hazlett, and Martin Styner, “Asymmetric bias in user guided segmentations of brain structures.,” NeuroImage, vol. 59, no. 2, pp. 1315–1323, Jan. 2012. |
| − | + | ||
|- | |- | ||
| − | | | [[Image: | + | | | [[Image:UNCShape_ShapeCorrespondence.png|200px]] |
| | | | | | ||
== [[Projects:LocalStatisticalAnalysisViaPermutationTests|Local Statistical Analysis via Permutation Tests]] == | == [[Projects:LocalStatisticalAnalysisViaPermutationTests|Local Statistical Analysis via Permutation Tests]] == | ||
| − | We have further developed a set of statistical testing methods that allow the analysis of local shape differences | + | We have further developed a set of statistical testing methods that allow the analysis of local shape differences via group differences tests as well interaction tests. Resulting significance maps (both raw and corrected for multiple comparisons) are easily visualized. Additional visualization of the group tests are provided via mean difference magnitude and vector maps, as well as maps of the group covariance information. Additional visualization of the interaction tests include Pearson and Spearman correlation maps. [[Projects:LocalStatisticalAnalysisViaPermutationTests|More...]] |
| − | <font color="red">'''New: '''</font> Paniagua B., Styner M., Macenko M., Pantazis D., Niethammer M, Local Shape Analysis using MANCOVA, Insight Journal, 2009 July-December, http://hdl.handle.net/10380/3124 | + | <font color="red">'''New: '''</font> User-friendly GUI interface and statistical result visualization via automatically generated Slicer MRML scenes |
| + | |||
| + | <font color="red">'''New: '''</font> Available on NITRC either [http://www.nitrc.org/projects/shape_mancova separately (ShapeAnalysisMANCOVA)] or as part of the [http://www.nitrc.org/projects/spharm-pdm SPHARM-PDM shape analysis package] | ||
| + | |||
| + | * Paniagua B., Styner M., Macenko M., Pantazis D., Niethammer M, Local Shape Analysis using MANCOVA, Insight Journal, 2009 July-December, http://hdl.handle.net/10380/3124 | ||
| − | |||
|- | |- | ||
| Line 70: | Line 154: | ||
In this project, we want to focus on the evaluation of medical image analysis methods for specific clinical applications in respect to development of evaluation methodology and the organization of venues promoting such comparison and validation studies. | In this project, we want to focus on the evaluation of medical image analysis methods for specific clinical applications in respect to development of evaluation methodology and the organization of venues promoting such comparison and validation studies. | ||
| − | <font color="red">'''New: '''</font> | + | <font color="red">'''New: '''</font> [[DTI_Tractography_Challenge_MICCAI_2012 | Wiki]] or [http://dti-challenge.org/ webpage] of DTI fiber tractography challengeat MICCAI 2012 |
| + | |||
| + | <font color="red">'''New: '''</font> DTI fiber tractography challenge at [[Events:_DTI_Tractography_Challenge_MICCAI_2011 |MICCAI 2011]] | ||
| + | |||
| + | |- | ||
| + | |||
| + | |||
| + | | | [[Image:UNCShape_CaudatePval_MICCAI06.png|200px]] | ||
| + | | | | ||
| + | |||
| + | == [[Projects:PopulationBasedCorrespondence|Population Based Correspondence]] == | ||
| + | |||
| + | We are developing methodology to automatically find dense point correspondences between a collection of polygonal genus 0 meshes. The advantage of this method is independence from indivisual templates, as well as enhanced modeling properties. The method is based on minimizing a cost function that describes the goodness of correspondence. Apart from a cost function derived from the description length of the model, we also employ a cost function working with arbitrary local features. We extended the original methods to use surface curvature measurements, which are independent to differences of object aligment. [[Projects:PopulationBasedCorrespondence|More...]] | ||
| + | |||
| + | * Styner M., Oguz I., Heimann T., Gerig G. Minimum description length with local geometry. Proceedings of the 5th IEEE International Symposium on Biomedical Imaging: From Nano to Macro 2008; 1283-1286 | ||
| + | * Software available as part of UNC Neurolib open source ([http://www.ia.unc.edu/dev website]) | ||
|- | |- | ||
| + | |||
|} | |} | ||
Latest revision as of 21:39, 11 December 2012
Home < Algorithm:UNCBack to NA-MIC Algorithms
Overview of UNC Algorithms (PI: Martin Styner)
At UNC, we are interested in a range of algorithms and solutions for the surface based analysis of brain structures and the cortex. We pioneered the use of spherical harmonics based shape analysis for comparing brain structures across objects. We has also worked on incorporating various data sources for correspondence computation on surfaces of different complexity (ranging from simple brain structures to the highly folded cortical surface). A current topic includes the use of diffusion tensor imaging for connectivity analysis in pathological settings. Finally, investigating quality control, validation and evaluation methodology is another important topic of our NA-MIC research.
UNC Projects
|
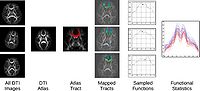
Atlas Based DTI Fiber Analysis FrameworkThis project aims to define an automatic framework for statistical comparison of fiber bundle diffusion properties between populations of diffusion weighted images. Quality control is performed on diffusion weighted images and DTI images are computed for each individual subject. The data is then either mapped into a prior atlas or an unbiased diffeomorphic DTI atlas is generated from all datasets. This creates a normalized coordinate system for all diffusion images in a study. Fiber tracts of interest are generated on this atlas, and then mapped back to the individual subjects. Diffusion properties along fiber tracts, such as fractional anisotropy (FA), are modeled as multivariate functions of arc length and gathered in spreadsheets for statistical analysis. More... New: Update versions on NITRC for DTIAtlasFiberAnalyzer and FiberViewerLight and DTIAtlasBuilder New: AR Verde, JB Berger, A Gupta, M Farzinfar, A Kaiser, VW Chanon, C Boettiger, C Goodlett, Y Shi, H Zhu, G Gerig, S Gouttard, C Vachet, M Styner, The UNC-Utah NA-MIC DTI framework: Atlas Based Fiber Tract Analysis with Application to a Study of Nicotine Smoking Addiction, accepted to SPIE Medical Imaging 2013 |
|
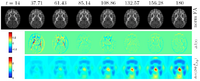
Longitudinal Atlas BuildingAs part of the longitudinal intra- and interpatient analysis theme within NA-MIC, we are working on a deformable, longitudinal DTI atlas method. Our longitudinal framework explicitly accounts for temporal dependencies via iterative subject-specific statistical growth modeling, and cross-sectional atlas-building. To effectively account for measurements sparse in time, a continuous-discrete statistical growth model is proposed incorporating also patient co-variatesMore... New: I. Csapo, B. Davis, Y. Shi, M. Sanchez, M. Styner, and M. Niethammer, Temporally-dependent image similarity measure for longitudinal analysis, presented at the WBIR'12: Proceedings of the 5th international conference on Biomedical Image Registration, 2012, vol. 7359, pp. 99–109. New: I. Csapo, B. Davis, Y. Shi, M. Sanchez, M. Styner, and M. Niethammer, “Longitudinal Image Registration with Non-uniform Appearance Change,” Medical Image Computing and Computer-Assisted Intervention – MICCAI 2012, vol. 7512, pp. 280–288, Aug. 2012. New: Y. Hong, S. Joshi, M. Sanchez, M. Styner, and M. Niethammer, Metamorphic Geodesic Regression, Medical Image Computing and Computer Assisted Interventions MICCAI 2012, vol. 7512, pp. 197–205, Aug. 2012. |
|
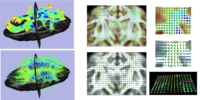
Diffusion Weighted Atlas Construction via model-based transformation and averaging of signalThis project investigated a method for model-based averaging of sets of diffusion weighted magnetic resonance images (DW-MRI) under space transformations (resulting for example from registration methods). A robust weighted least squares method is developed. Synthetic validation experiments show the improvement of the proposed estimation method in comparison to standard least squares estimation. The developed method is applied to construct an atlas of {\it diffusion weighted images} for a set of macaques, allowing for a more flexible representation of average diffusion information compared to standard diffusion tensor atlases. More...
New: M. Niethammer, Y. Shi, S. Benzaid, M. Sanchez, and M. Styner. Robust model-based transformation and averaging of diffusion weighted images applied to diffusion weighted atlas construction. MICCAI, Workshop on Computational Diffusion MRI, 2010. |

|
Diffusion Imaging based ConnectivityThis project focuses on connectivity measurements derived from diffusion imaging datasets in order to better understand cortical and subcortical white matter connectivity. Our research employs a novel, multi-directional graph propagation method that performs a fully deterministic, efficient and stable connectivity computation. The method handles crossing fibers and deals well with multiple seed regions. In addition to the analysis of these connectivity measures in describing brain pathology, they can also be used as scalar maps for use in DTI registration. More... New: I. Oguz, A. Boucharin, W. Lu, C. Vachet, F. Budin, Y. Shi, and M. Styner, A Minimum Cost Approach to Connectivity from Orientation Distribution Functions via Efficient Multi-directional Graph Propagation, Proceedings of CDMRI workshop, MICCAI 2012, pp. 210–221, Dec. 2012. New: Alexis Boucharin, Ipek Oguz, Clement Vachet, Yundi Shi, Mar Sanchez, Martin Styner. Efficient, graph-based white matter connectivity from orientation distribution functions via multi-directional graph propagation. Medical Imaging 2011: Image Processing (2011) vol. 7962 (1) pp. 79620S |
|
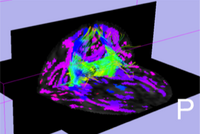
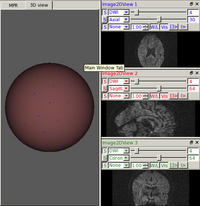
DWI and DTI Quality ControlDWI data suffers from inherent low SNR, overall long scanning time of multiple directional encoding with correspondingly large risk to encounter several kinds of artifacts. These artifacts can be too severe for a correct and stable estimation of the diffusion tensor. Thus, a quality control (QC) procedure is absolutely necessary for DTI studies. We are developing a framework for automatic DWI and DTI quality assessment and correction. We developed a tool called DTIPrep which pipelines the QC steps with designated protocol use and report generation. More... New: Mahshid Farzinfar, Yinpeng Li, Audrey Verde, Ipek Oguz, Guido Gerig, Martin A. Styner, DTI quality control assessment via error estimation from Monte Carlo simulation, to be presented at SPIE Medical Imaging 2013. New: M. Farzinfar, C. Dietrich, R. G. Smith, Y. Li, A. Gupta, Z. Liu, and M. Styner, “Entropy based DTI quality control via regional orientation distribution,” Biomedical Imaging (ISBI), 2012 9th IEEE International Symposium on, pp. 22–25, 2012. New: DTIPrep new version on NITRC , including visual quality control support |
|
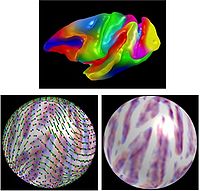
Cortical Correspondence using Sulcal CurvesIn this project, we compute group wise cortical correspondence on populations, using information from sulcal curves, as well as a novel way to encode deformations on the sphere. This framework is then applied to data from human and non-human primate cortical thickness studies. More... New: Ilwoo Lyu, Sun Hyung Kim, Joon-Kyung Seong, Soongsil Univ., Sang Wook Yoo, Alan Evans, Yundi Shi, Mar Sanchez, Marc Niethammer, Martin A. Styner, Cortical correspondence via sulcal curve-constrained spherical registration with application to Macaque studies, to be presented at SPIE Medical Imaging 2013 |
|
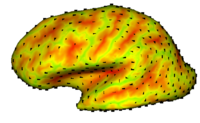
Cortical Correspondence using Particle SystemIn this project, we want to compute cortical correspondence on populations, using various features such as cortical structure, DTI connectivity, vascular structure, and functional data (fMRI). This presents a challenge because of the highly convoluted surface of the cortex, as well as because of the different properties of the data features we want to incorporate together. This correspondence method has been included in our NAMIC cortical thickness framework GAMBIT More... New: Vachet, C., Hazlett, H., Niethammer, M., Oguz, I., Cates, J., Whitaker, R., Piven, J., Styner, M., “Group-wise automatic mesh-based analysis of cortical thickness“. Medical Imaging 2011: Image Processing (2011) vol. 7962 (1) pp. 796227 1 - 10 New: Lee J, Ehlers C, Crews F, Niethammer M, Budin F, Paniagua B, Sulik K, Johns J, Styner M, Oguz I. Automatic cortical thickness analysis on rodent brain. Medical Imaging 2011: Image Processing (2011) vol. 7962 (1) pp. 796248 |

|
UNC-Utah Shape Analysis FrameworkThe UNC shape analysis is based on an analysis framework of objects with spherical topology, described mainly by sampled spherical harmonics SPHARM-PDM. The input of the shape analysis framework is a set of binary segmentations of a single brain structure, such as the hippocampus or caudate. These segmentations are converted into a shape description (SPHARM-PDM) with correspondence and tested via statistical point-wise analysis. Additionally, the SPHARM correspondences can be improved with Entropy-based particle systems, by using an integration module recently added to the pipeline. More...
New: N. Solowij, M. Walterfang, D. I. Lubman, S. Whittle, V. Lorenzetti, M. Styner, D. Velakoulis, C. Pantelis, and M. Yücel, “Alteration to hippocampal shape in cannabis users with and without schizophrenia.,” Schizophr Res, Nov. 2012. New: O. Lindberg, M. Walterfang, J. C. L. Looi, N. Malykhin, P. Ostberg, B. Zandbelt, M. Styner, B. Paniagua, D. Velakoulis, E. Orndahl, and L.-O. Wahlund, “Hippocampal Shape Analysis in Alzheimer's Disease and Frontotemporal Lobar Degeneration Subtypes.,” J Alzheimers Dis, Mar. 2012. New: D. Ong, M. A. Walterfang, G. Malhi, M. Styner, D. Velakoulis, and C. Pantelis, “Shape Alteration in the Caudate Nucleus in Individuals With Bipolar Affective Disorder Disorder.,” Aust N Z J Psychiatry, Feb. 2012. New: Beatriz Paniagua, Omri Emodi, Jonathan Hill, James Fishbaugh, Luiz Pimenta, Stephen Aylward, Andinet Enquobahrie, Guido Gerig, John Gilmore, John van Aalst, Martin A. Styner, 3D of brain shape and volume after cranial vault remodeling surgery for craniosynostosis correction in infants, to be published in SPIE Medical Imaging 2013 New: Beatriz Paniagua, Amanda Lyall, Jean-Baptiste Berger, Clement Vachet, Robert Hamer, Sandra Woolson, Weili Lin, John Gilmore, Martin A. Styner, Lateral ventricle morphology analysis via mean latitude axis, to be published in SPIE Medical Imaging 2013 New: Eric Maltbie, Kshamta Bhatt, Beatriz Paniagua, Rachel Smith, Michael Graves, Matthew Mosconi, Sarah Peterson, Scott White, Joseph Blocher, Mohammed El-Sayed, Heather Hazlett, and Martin Styner, “Asymmetric bias in user guided segmentations of brain structures.,” NeuroImage, vol. 59, no. 2, pp. 1315–1323, Jan. 2012.
|

|
Local Statistical Analysis via Permutation TestsWe have further developed a set of statistical testing methods that allow the analysis of local shape differences via group differences tests as well interaction tests. Resulting significance maps (both raw and corrected for multiple comparisons) are easily visualized. Additional visualization of the group tests are provided via mean difference magnitude and vector maps, as well as maps of the group covariance information. Additional visualization of the interaction tests include Pearson and Spearman correlation maps. More... New: User-friendly GUI interface and statistical result visualization via automatically generated Slicer MRML scenes New: Available on NITRC either separately (ShapeAnalysisMANCOVA) or as part of the SPHARM-PDM shape analysis package
|

|
Evaluation and Comparison of Medical Image Analysis MethodsIn this project, we want to focus on the evaluation of medical image analysis methods for specific clinical applications in respect to development of evaluation methodology and the organization of venues promoting such comparison and validation studies. New: Wiki or webpage of DTI fiber tractography challengeat MICCAI 2012 New: DTI fiber tractography challenge at MICCAI 2011 |
|
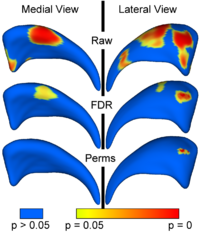
Population Based CorrespondenceWe are developing methodology to automatically find dense point correspondences between a collection of polygonal genus 0 meshes. The advantage of this method is independence from indivisual templates, as well as enhanced modeling properties. The method is based on minimizing a cost function that describes the goodness of correspondence. Apart from a cost function derived from the description length of the model, we also employ a cost function working with arbitrary local features. We extended the original methods to use surface curvature measurements, which are independent to differences of object aligment. More...
|